Prerequisites: Vector Length / Norm (ch128), Geometric Interpretation (ch122) You will learn:
How distance between two vectors is computed from their difference
The connection between norm and metric
Applications: nearest neighbor search, clustering, similarity
Environment: Python 3.x, numpy, matplotlib
1. Concept¶
The distance between two vectors u and v is the norm of their difference:
This generalizes the familiar 2D distance formula to any number of dimensions, and to any choice of norm. The Euclidean distance (L2) is the most common, but L1 and L∞ distances each have natural applications.
Common misconception: The distance between two vectors is not np.dot(u - v, u - v) — that is the squared distance. Always take the square root for the actual distance.
2. Intuition & Mental Models¶
Geometric model: The distance from u to v is the length of the arrow from v to u (the vector u − v). This arrow is the displacement vector discussed in ch122.
Metric model: A distance function is a metric if it satisfies: non-negativity, symmetry (), identity (), and triangle inequality (). Every norm induces a valid metric by .
Recall from ch128 (Vector Length): is the norm. Distance is — the norm of the displacement.
3. Visualization & Implementation¶
# --- Visualization: Distance as the norm of the difference ---
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-v0_8-whitegrid')
u = np.array([1.0, 4.0])
v = np.array([4.0, 1.0])
diff = u - v
d_l2 = np.linalg.norm(diff, 2)
d_l1 = np.linalg.norm(diff, 1)
fig, ax = plt.subplots(figsize=(6, 6))
ax.plot(*u, 'bo', markersize=10, label='u')
ax.plot(*v, 'ro', markersize=10, label='v')
ax.annotate('', xy=u, xytext=v,
arrowprops=dict(arrowstyle='->', color='navy', lw=2.5))
ax.text(2.4, 2.8, f'd_L2={d_l2:.3f}', fontsize=11, color='navy')
ax.plot([v[0], u[0], u[0]], [v[1], v[1], u[1]], 'k--', lw=1, label=f'd_L1={d_l1:.2f}')
ax.set_xlim(0, 6); ax.set_ylim(0, 6)
ax.set_aspect('equal')
ax.set_title('Distance as ||u - v||')
ax.set_xlabel('x'); ax.set_ylabel('y')
ax.legend()
plt.tight_layout(); plt.show()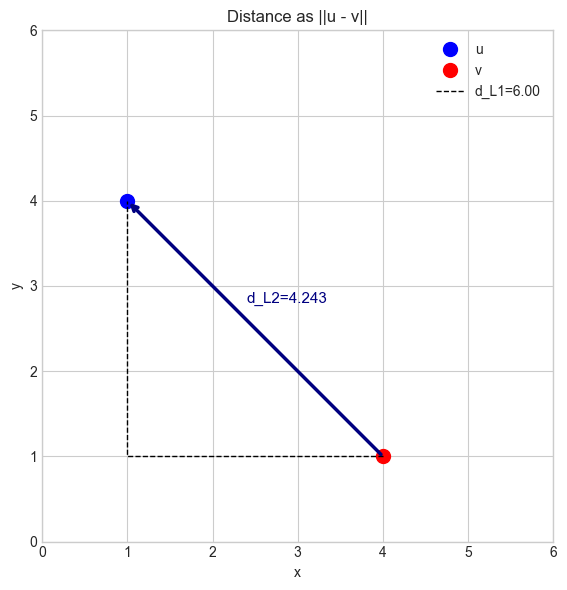
# --- Implementation: distance functions ---
import numpy as np
def euclidean_distance(u, v):
"""L2 distance between u and v."""
return np.linalg.norm(np.asarray(u, float) - np.asarray(v, float))
def manhattan_distance(u, v):
"""L1 distance between u and v."""
return np.linalg.norm(np.asarray(u, float) - np.asarray(v, float), 1)
def pairwise_distances(X, p=2):
"""
Compute all pairwise Lp distances between rows of X.
Args:
X: ndarray shape (n, d)
p: norm order
Returns:
D: ndarray shape (n, n) — D[i,j] = dist(X[i], X[j])
"""
n = X.shape[0]
D = np.zeros((n, n))
for i in range(n):
for j in range(i+1, n):
d = np.linalg.norm(X[i] - X[j], p)
D[i, j] = D[j, i] = d
return D
# Test
u = np.array([1.0, 4.0])
v = np.array([4.0, 1.0])
print("Euclidean:", euclidean_distance(u, v))
print("Manhattan:", manhattan_distance(u, v))
X = np.array([[0,0],[3,4],[1,1]], dtype=float)
D = pairwise_distances(X)
print("\nPairwise L2 distances:\n", D)Euclidean: 4.242640687119285
Manhattan: 6.0
Pairwise L2 distances:
[[0. 5. 1.41421356]
[5. 0. 3.60555128]
[1.41421356 3.60555128 0. ]]
4. Experiments¶
# --- Experiment: Curse of dimensionality — distances concentrate ---
# Hypothesis: in high dimensions, all pairwise distances converge to the same value.
# Try changing: DIM
import numpy as np
N = 500 # number of points
np.random.seed(0)
for DIM in [2, 10, 50, 200, 1000]:
X = np.random.randn(N, DIM)
# compute a sample of pairwise distances
idx1 = np.random.randint(0, N, 1000)
idx2 = np.random.randint(0, N, 1000)
dists = np.linalg.norm(X[idx1] - X[idx2], axis=1)
print(f"dim={DIM:5d} mean={dists.mean():.3f} std={dists.std():.3f} cv={dists.std()/dists.mean():.4f}")dim= 2 mean=1.714 std=0.918 cv=0.5357
dim= 10 mean=4.434 std=1.018 cv=0.2295
dim= 50 mean=9.923 std=1.055 cv=0.1063
dim= 200 mean=19.797 std=1.581 cv=0.0798
dim= 1000 mean=44.690 std=0.976 cv=0.0218
5. Exercises¶
Easy 1. Compute L2 distance between (1,2,3) and (4,6,3). (Expected: 5)
Medium 1. Given 5 points in 3D, find the pair with minimum L2 distance. Do the same with L1 distance. Are the closest pairs the same?
Hard. Implement an O(n²) brute-force all-pairs distance matrix for n=500 points in 10D. Time it. Then implement the same using NumPy broadcasting without explicit loops. Report the speedup.
6. Summary & Connections¶
Distance between vectors: .
Every norm defines a valid metric (distance function).
In high dimensions, L2 distances concentrate — motivation for cosine similarity (ch132).
Backward connection: Generalizes the 2D distance formula from ch094 to arbitrary dimension and norm choice.
Forward connections:
This will reappear in ch135 — Orthogonality, where two vectors at 90° have a specific distance relationship.
This will reappear in ch175 — Linear Regression, where residuals are distances in the output space.