(Solves the hyperparameter selection problem from ch283; generalizes bootstrap from ch275)
1. The Problem with a Single Train/Test Split¶
A single split wastes data (the test set can’t train) and produces a high-variance estimate of test error (it depends on which random split you got). Cross-validation uses the data more efficiently.
2. K-Fold Cross Validation¶
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import Ridge
rng = np.random.default_rng(0)
def kfold_indices(n: int, k: int, rng) -> list[tuple[np.ndarray, np.ndarray]]:
"""Returns list of (train_idx, val_idx) tuples for k-fold CV."""
shuffled = rng.permutation(n)
fold_size = n // k
folds = []
for i in range(k):
val_idx = shuffled[i*fold_size:(i+1)*fold_size]
train_idx = np.concatenate([shuffled[:i*fold_size], shuffled[(i+1)*fold_size:]])
folds.append((train_idx, val_idx))
return folds
def cross_validate(
X: np.ndarray,
y: np.ndarray,
model_fn, # fn(X_tr, y_tr, X_val) -> y_pred_val
metric_fn, # fn(y_true, y_pred) -> float
k: int = 5,
rng = None,
) -> tuple[float, float]:
"""Returns (mean_score, std_score)."""
if rng is None: rng = np.random.default_rng()
folds = kfold_indices(len(X), k, rng)
scores = []
for train_idx, val_idx in folds:
y_pred = model_fn(X[train_idx], y[train_idx], X[val_idx])
scores.append(metric_fn(y[val_idx], y_pred))
return np.mean(scores), np.std(scores)
# Generate dataset
n = 200
X_raw = rng.normal(0, 1, (n, 5))
y_raw = 2 + X_raw @ np.array([1.5, -2.0, 0.5, 0, 0]) + rng.normal(0, 1.5, n)
# Ridge regression with different lambdas
lambdas = np.logspace(-4, 3, 30)
cv_means, cv_stds = [], []
def mse(y_true, y_pred): return np.mean((y_true - y_pred)**2)
for lam in lambdas:
def model(X_tr, y_tr, X_val, _lam=lam):
# Use ridge_regression from ch283 — here using sklearn to avoid copying
m = Ridge(alpha=_lam, fit_intercept=True)
m.fit(X_tr, y_tr)
return m.predict(X_val)
mu, sigma = cross_validate(X_raw, y_raw, model, mse, k=5, rng=rng)
cv_means.append(mu)
cv_stds.append(sigma)
cv_means = np.array(cv_means)
cv_stds = np.array(cv_stds)
best_lam = lambdas[np.argmin(cv_means)]
fig, ax = plt.subplots(figsize=(9, 4))
ax.semilogx(lambdas, cv_means, 'b-o', ms=4, label='CV MSE')
ax.fill_between(lambdas, cv_means - cv_stds, cv_means + cv_stds, alpha=0.2)
ax.axvline(best_lam, color='red', ls='--', label=f'Best λ={best_lam:.4f}')
ax.set_xlabel('λ (log scale)')
ax.set_ylabel('5-Fold CV MSE')
ax.set_title('Cross-Validation for Hyperparameter Selection')
ax.legend()
plt.tight_layout()
plt.show()
print(f"Best λ: {best_lam:.4f}")
print(f"CV MSE at best λ: {cv_means.min():.4f} ± {cv_stds[np.argmin(cv_means)]:.4f}")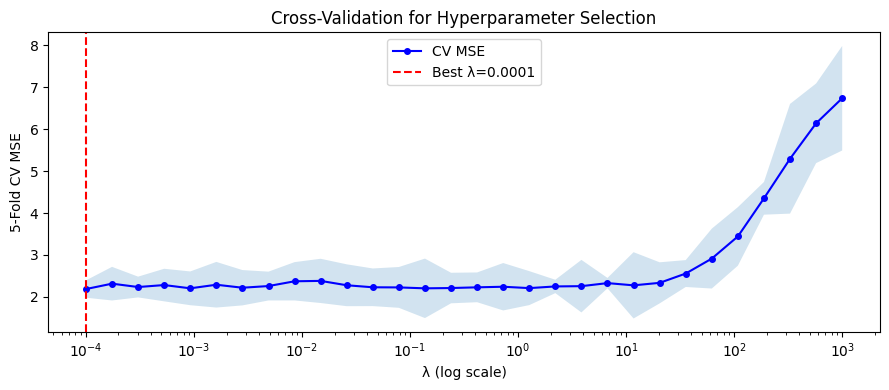
Best λ: 0.0001
CV MSE at best λ: 2.1859 ± 0.2066
3. Leave-One-Out and Stratified K-Fold¶
def stratified_kfold_indices(
y: np.ndarray, k: int, rng
) -> list[tuple[np.ndarray, np.ndarray]]:
"""
For binary classification: ensure each fold has the same class ratio.
"""
classes = np.unique(y)
class_indices = [rng.permutation(np.where(y == c)[0]) for c in classes]
folds_per_class = [np.array_split(idx, k) for idx in class_indices]
folds = []
for i in range(k):
val_idx = np.concatenate([f[i] for f in folds_per_class])
train_idx = np.concatenate([
np.concatenate([f[j] for j in range(k) if j != i])
for f in folds_per_class
])
folds.append((train_idx, val_idx))
return folds
# Verify class balance in stratified vs regular k-fold
y_imbalanced = np.array([1]*50 + [0]*150)
true_ratio = y_imbalanced.mean()
regular_ratios = []
stratified_ratios = []
for train_idx, val_idx in kfold_indices(len(y_imbalanced), 5, rng):
regular_ratios.append(y_imbalanced[val_idx].mean())
for train_idx, val_idx in stratified_kfold_indices(y_imbalanced, 5, rng):
stratified_ratios.append(y_imbalanced[val_idx].mean())
print(f"True positive ratio: {true_ratio:.3f}")
print(f"Regular k-fold fold ratios: {[f'{r:.3f}' for r in regular_ratios]}")
print(f"Stratified k-fold fold ratios: {[f'{r:.3f}' for r in stratified_ratios]}")True positive ratio: 0.250
Regular k-fold fold ratios: ['0.325', '0.125', '0.225', '0.250', '0.325']
Stratified k-fold fold ratios: ['0.250', '0.250', '0.250', '0.250', '0.250']
4. What Comes Next¶
Cross-validation is the standard evaluation procedure for any ML system. ch285 — A/B Testing extends hypothesis testing to the setting where you compare two systems on live traffic, with the same concern for multiple comparisons that appeared in ch278. The full model selection workflow — CV for hyperparameters, held-out test set for final evaluation — is implemented in the project chapters ch298–299.