(Directly extends ch288 — Entropy; appears in every generative model loss function)
1. The Definition¶
The Kullback-Leibler divergence from to measures how much information is lost when is used to approximate :
Properties:
always (Gibbs’ inequality)
iff almost everywhere
Not symmetric: in general
Not a distance metric (triangle inequality fails)
Connection to cross-entropy and entropy:
Minimizing cross-entropy ↔ minimizing KL divergence (since is fixed).
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
def kl_divergence_discrete(
p: np.ndarray,
q: np.ndarray,
eps: float = 1e-15,
) -> float:
"""D_KL(P||Q) in nats. Assumes p,q sum to 1."""
p = np.asarray(p, dtype=float)
q = np.asarray(q, dtype=float)
mask = p > 0
return np.sum(p[mask] * np.log(p[mask] / np.clip(q[mask], eps, 1.0)))
def entropy_discrete(p: np.ndarray) -> float:
p = np.asarray(p, dtype=float)
mask = p > 0
return -np.sum(p[mask] * np.log(p[mask]))
def cross_entropy_discrete(p: np.ndarray, q: np.ndarray) -> float:
return entropy_discrete(p) + kl_divergence_discrete(p, q)
# Verify decomposition
p_true = np.array([0.5, 0.3, 0.2])
q_pred = np.array([0.3, 0.4, 0.3])
h_p = entropy_discrete(p_true)
kl = kl_divergence_discrete(p_true, q_pred)
ce = cross_entropy_discrete(p_true, q_pred)
ce_dir = -np.sum(p_true * np.log(q_pred))
print(f"H(P): {h_p:.6f}")
print(f"KL(P||Q): {kl:.6f}")
print(f"H(P,Q): {ce:.6f}")
print(f"Direct CE: {ce_dir:.6f}")
print(f"H(P)+KL=CE: {np.isclose(h_p + kl, ce)}")
print()
print(f"D_KL(P||Q) = {kl:.4f}")
print(f"D_KL(Q||P) = {kl_divergence_discrete(q_pred, p_true):.4f} (asymmetric!)")H(P): 1.029653
KL(P||Q): 0.088015
H(P,Q): 1.117668
Direct CE: 1.117668
H(P)+KL=CE: True
D_KL(P||Q) = 0.0880
D_KL(Q||P) = 0.0835 (asymmetric!)
2. Forward vs Reverse KL¶
The asymmetry has practical consequences:
Forward KL : minimizing forces to cover all modes of (zero-avoiding)
Reverse KL : minimizing forces to concentrate on one mode of (mean-seeking)
This determines the behavior of variational inference methods.
# Approximate a bimodal distribution with a Gaussian
# Forward KL: spread to cover both modes
# Reverse KL: collapse to one mode
x = np.linspace(-5, 5, 1000)
dx = x[1] - x[0]
# True distribution P: bimodal
p_true_cont = 0.5 * stats.norm.pdf(x, -2, 0.6) + 0.5 * stats.norm.pdf(x, 2, 0.6)
p_true_cont /= p_true_cont.sum() * dx # normalize
# Search for Gaussian Q that minimizes forward vs reverse KL
def gaussian_pdf(x, mu, sigma):
return stats.norm.pdf(x, mu, sigma)
def forward_kl(mu, sigma, p_true, x, dx):
"""D_KL(P||Q) for Gaussian Q."""
q = gaussian_pdf(x, mu, sigma)
q = np.clip(q, 1e-15, None)
p = np.clip(p_true, 1e-15, None)
return np.sum(p * np.log(p / q)) * dx
def reverse_kl(mu, sigma, p_true, x, dx):
"""D_KL(Q||P) for Gaussian Q."""
q = gaussian_pdf(x, mu, sigma)
q = np.clip(q, 1e-15, None)
p = np.clip(p_true, 1e-15, None)
return np.sum(q * np.log(q / p)) * dx
# Evaluate over a grid of (mu, sigma) and find minimizers
mus = np.linspace(-3, 3, 40)
sigmas = np.linspace(0.3, 3.0, 40)
fwd_losses = np.array([
[forward_kl(m, s, p_true_cont, x, dx) for s in sigmas] for m in mus
])
rev_losses = np.array([
[reverse_kl(m, s, p_true_cont, x, dx) for s in sigmas] for m in mus
])
best_fwd = np.unravel_index(fwd_losses.argmin(), fwd_losses.shape)
best_rev = np.unravel_index(rev_losses.argmin(), rev_losses.shape)
mu_fwd, sig_fwd = mus[best_fwd[0]], sigmas[best_fwd[1]]
mu_rev, sig_rev = mus[best_rev[0]], sigmas[best_rev[1]]
fig, ax = plt.subplots(figsize=(9, 4))
ax.plot(x, p_true_cont, 'k-', lw=2.5, label='True P (bimodal)')
ax.plot(x, gaussian_pdf(x, mu_fwd, sig_fwd), 'b-', lw=2,
label=f'Min Forward KL: μ={mu_fwd:.2f}, σ={sig_fwd:.2f}')
ax.plot(x, gaussian_pdf(x, mu_rev, sig_rev), 'r-', lw=2,
label=f'Min Reverse KL: μ={mu_rev:.2f}, σ={sig_rev:.2f}')
ax.set_title('Forward KL → covers both modes | Reverse KL → one mode')
ax.set_xlabel('x'); ax.legend(fontsize=9)
plt.tight_layout()
plt.show()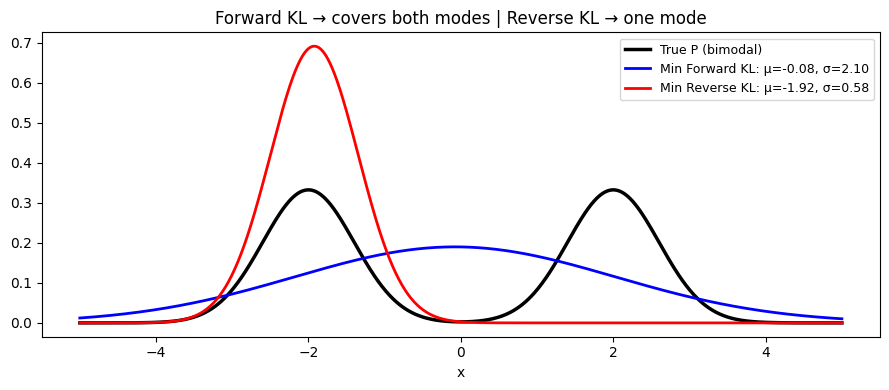
3. What Comes Next¶
ch290 — Mutual Information combines entropy and KL divergence to measure how much two variables share. It is the non-linear, model-free generalization of correlation — applicable wherever Pearson’s is not.