(Applies SVD from ch173 and PCA intuition from ch174; uses covariance from ch273)
1. The Curse of Dimensionality¶
In high-dimensional spaces, every point is far from every other point. Data becomes sparse. Distances lose meaning. Models require exponentially more samples to generalize.
Dimensionality reduction finds a lower-dimensional representation that preserves the structure relevant to your task.
2. PCA from Scratch¶
import numpy as np
import matplotlib.pyplot as plt
from sklearn.decomposition import PCA as sklearn_PCA
from sklearn.datasets import load_digits
class PCA:
"""
Principal Component Analysis via eigendecomposition of the covariance matrix.
Numerically, uses SVD for stability (equivalent, more robust).
"""
def __init__(self, n_components: int):
self.n_components = n_components
self.components_ = None # principal components (rows)
self.explained_variance_ratio_ = None
self.mean_ = None
def fit(self, X: np.ndarray) -> 'PCA':
self.mean_ = X.mean(axis=0)
X_c = X - self.mean_
# SVD: X_c = U @ S @ Vt
# Principal components are rows of Vt
_, S, Vt = np.linalg.svd(X_c, full_matrices=False)
self.components_ = Vt[:self.n_components]
# Explained variance from singular values
explained_var = (S**2) / (len(X) - 1)
self.explained_variance_ratio_ = (
explained_var[:self.n_components] / explained_var.sum()
)
self._singular_values = S
return self
def transform(self, X: np.ndarray) -> np.ndarray:
return (X - self.mean_) @ self.components_.T
def fit_transform(self, X: np.ndarray) -> np.ndarray:
return self.fit(X).transform(X)
def inverse_transform(self, X_reduced: np.ndarray) -> np.ndarray:
return X_reduced @ self.components_ + self.mean_
# Generate 3D data with intrinsic 2D structure
rng = np.random.default_rng(42)
n = 300
t = rng.uniform(0, 2*np.pi, n)
X_3d = np.column_stack([
np.cos(t) + rng.normal(0, 0.05, n),
np.sin(t) + rng.normal(0, 0.05, n),
0.2 * t + rng.normal(0, 0.05, n),
])
pca = PCA(n_components=2)
X_2d = pca.fit_transform(X_3d)
# Validate against sklearn
pca_sk = sklearn_PCA(n_components=2)
X_2d_sk = pca_sk.fit_transform(X_3d)
print("Explained variance ratio (manual):",
np.round(pca.explained_variance_ratio_, 4))
print("Explained variance ratio (sklearn):",
np.round(pca_sk.explained_variance_ratio_, 4))
print("Match (up to sign):",
np.allclose(np.abs(X_2d), np.abs(X_2d_sk), atol=1e-8))
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
ax = axes[0]
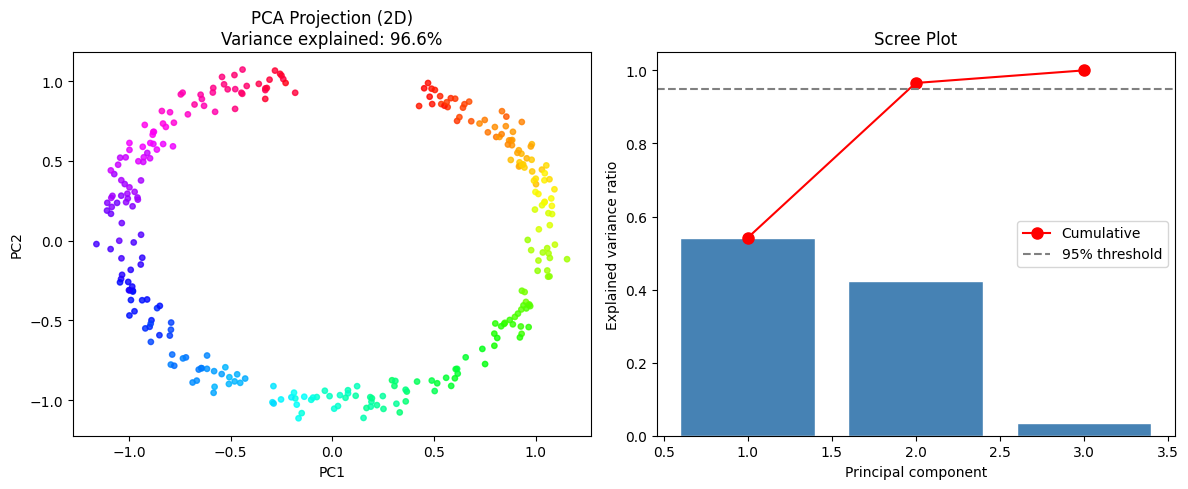
ax.scatter(X_2d[:, 0], X_2d[:, 1], c=t, cmap='hsv', s=15, alpha=0.8)
ax.set_title(f'PCA Projection (2D)\n'
f'Variance explained: {pca.explained_variance_ratio_.sum():.1%}')
ax.set_xlabel('PC1'); ax.set_ylabel('PC2')
# Scree plot
pca_full = PCA(n_components=3).fit(X_3d)
ax = axes[1]
explained = (pca_full._singular_values**2) / (pca_full._singular_values**2).sum()
ax.bar(range(1, 4), explained, color='steelblue', edgecolor='white')
ax.plot(range(1, 4), np.cumsum(explained), 'ro-', ms=8, label='Cumulative')
ax.axhline(0.95, color='gray', ls='--', label='95% threshold')
ax.set_xlabel('Principal component')
ax.set_ylabel('Explained variance ratio')
ax.set_title('Scree Plot')
ax.legend()
plt.tight_layout()
plt.show()Explained variance ratio (manual): [0.5416 0.4239]
Explained variance ratio (sklearn): [0.5416 0.4239]
Match (up to sign): True
3. PCA for Reconstruction and Compression¶
# Apply PCA to handwritten digits
digits = load_digits()
X_digits = digits.data # 1797 x 64
y_digits = digits.target
pca_dig = PCA(n_components=40)
X_reduced = pca_dig.fit_transform(X_digits)
X_reconstructed = pca_dig.inverse_transform(X_reduced)
# Show compression effect
n_comps = [2, 5, 10, 20, 40, 64]
rec_errors = []
for k in n_comps:
pk = PCA(n_components=min(k, 64))
Xr = pk.fit_transform(X_digits)
Xrec = pk.inverse_transform(Xr)
err = np.mean((X_digits - Xrec)**2)
rec_errors.append(err)
fig, axes = plt.subplots(1, 2, figsize=(13, 4))
# Reconstruction at different k
ax = axes[0]
ax.plot(n_comps, rec_errors, 'o-', color='steelblue', lw=2)
ax.set_xlabel('Number of principal components')
ax.set_ylabel('Reconstruction MSE')
ax.set_title('Reconstruction Error vs Compression Level')
# 2D projection of digits
pca2 = PCA(n_components=2)
X_2d_digits = pca2.fit_transform(X_digits)
ax = axes[1]
for digit in range(10):
mask = y_digits == digit
ax.scatter(X_2d_digits[mask, 0], X_2d_digits[mask, 1],
s=5, alpha=0.6, label=str(digit))
ax.set_title('Digits in PCA 2D Space\n'
f'Variance: {pca2.explained_variance_ratio_.sum():.1%}')
ax.legend(fontsize=6, ncol=2)
plt.tight_layout()
plt.show()
print(f"Original: {X_digits.shape} = {X_digits.nbytes // 1024} KB")
print(f"After PCA (k=40): {X_reduced.shape} = {X_reduced.nbytes // 1024} KB")
print(f"Compression ratio: {X_digits.nbytes / X_reduced.nbytes:.1f}x")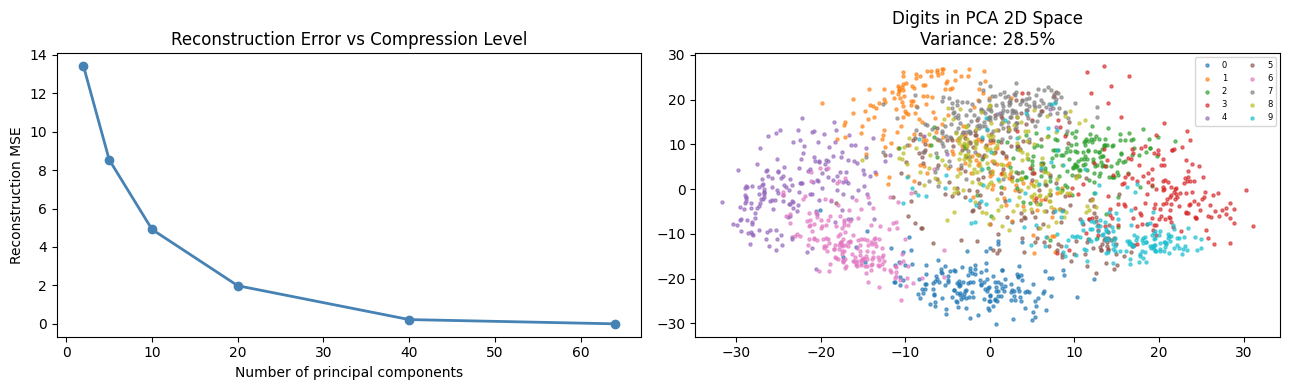
Original: (1797, 64) = 898 KB
After PCA (k=40): (1797, 40) = 561 KB
Compression ratio: 1.6x
4. t-SNE: Non-Linear Dimensionality Reduction (Concept)¶
# t-SNE is non-linear and preserves local structure.
# Full implementation is complex; we show the core idea and use sklearn.
from sklearn.manifold import TSNE
tsne = TSNE(n_components=2, perplexity=30, random_state=42)
X_tsne = tsne.fit_transform(X_digits[:500]) # subset for speed
y_sub = y_digits[:500]
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
for ax, (X_2d, title) in zip(axes, [
(pca2.transform(X_digits[:500]), 'PCA (linear)'),
(X_tsne, 't-SNE (non-linear)'),
]):
for digit in range(10):
mask = y_sub == digit
ax.scatter(X_2d[mask, 0], X_2d[mask, 1], s=8, alpha=0.7, label=str(digit))
ax.set_title(title)
ax.legend(fontsize=7, ncol=2)
plt.suptitle('PCA vs t-SNE on Digits', fontsize=11)
plt.tight_layout()
plt.show()
print("PCA: preserves global structure, linear, fast, deterministic.")
print("t-SNE: preserves local clusters, non-linear, slow, non-deterministic.")
print("Use PCA for compression/preprocessing; t-SNE for visualization.")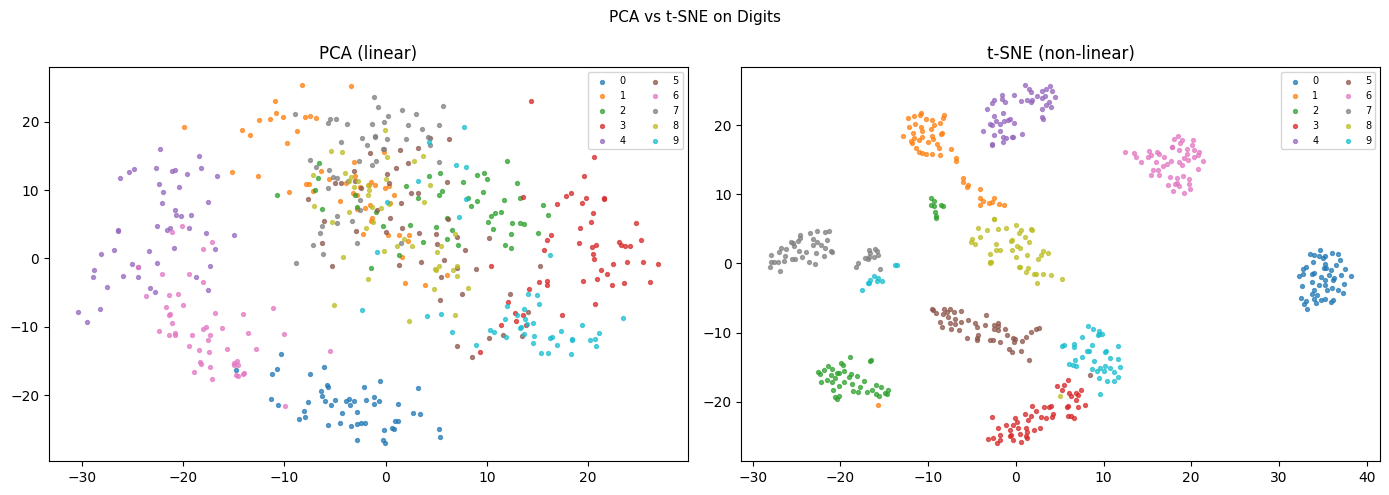
PCA: preserves global structure, linear, fast, deterministic.
t-SNE: preserves local clusters, non-linear, slow, non-deterministic.
Use PCA for compression/preprocessing; t-SNE for visualization.
5. What Comes Next¶
PCA is unsupervised — it finds directions of maximum variance without using labels. ch293 — Clustering is also unsupervised and finds group structure in data. ch294 — Classification is supervised and uses labels to find decision boundaries. The 2D PCA projections here make the class structure visible — a direct preview of what classifiers will formalize.