This chapter draws explicitly on all nine Parts. Every component traces to a specific prior chapter. Nothing is imported that has not been derived.
The System We Are Building¶
A complete document classification system:
Data — synthetic text feature vectors with noisy labels
Cleaning — missing value detection, outlier handling
Features — TF-IDF-style embeddings, normalization, dimensionality reduction
Model — neural network trained with backpropagation and Adam
Evaluation — cross-validation, ROC, calibration, confidence intervals
Information — entropy and mutual information of predictions
Deployment — streaming inference, uncertainty quantification
Each section states which Part(s) it draws from.
Part I Bridge: Mathematical Thinking¶
Ch001–020: abstraction, modeling, notation
The entire system is a mathematical model:
where is the feature space and is the -simplex (the set of valid probability distributions over classes). Every design decision is a choice within this mathematical framework.
import numpy as np
import matplotlib.pyplot as plt
from scipy import stats
from sklearn.datasets import fetch_20newsgroups
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.decomposition import TruncatedSVD
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import (
roc_auc_score, accuracy_score, classification_report
)
from sklearn.preprocessing import label_binarize
rng = np.random.default_rng(42)
print('All imports successful.')
print()
print('System: Document Classification — 4 newsgroup categories')
print('Input: raw text documents')
print('Output: probability distribution over 4 classes + uncertainty estimate')All imports successful.
System: Document Classification — 4 newsgroup categories
Input: raw text documents
Output: probability distribution over 4 classes + uncertainty estimate
Part II + III Bridge: Numbers, Functions, and Feature Representation¶
Ch021–090: numbers, logarithms (ch043), functions as programs (ch052)
TF-IDF is a function that maps raw text to a vector of real numbers. The IDF weight is a logarithm:
This is exponential decay of term weight (ch041 — Exponential Growth) applied in reverse: common words get log-compressed weights.
# Load 20 Newsgroups — 4 categories
categories = [
'sci.med', 'sci.space',
'rec.sport.hockey', 'rec.sport.baseball'
]
newsgroups = fetch_20newsgroups(
subset='all', categories=categories,
remove=('headers', 'footers', 'quotes'),
random_state=42
)
X_text = newsgroups.data
y_raw = newsgroups.target
labels = newsgroups.target_names
print(f'Dataset: {len(X_text)} documents, {len(labels)} classes')
for i, lbl in enumerate(labels):
print(f' Class {i}: {lbl} — {(y_raw==i).sum()} docs')
# TF-IDF vectorization
tfidf = TfidfVectorizer(
max_features=10_000,
min_df=3,
max_df=0.95,
sublinear_tf=True, # use log(1+tf) instead of raw tf
ngram_range=(1, 2),
)
X_tfidf = tfidf.fit_transform(X_text) # sparse matrix (n_docs, 10000)
print(f'\nTF-IDF matrix: {X_tfidf.shape}, density: {X_tfidf.nnz/np.prod(X_tfidf.shape):.4f}')Dataset: 3970 documents, 4 classes
Class 0: rec.sport.baseball — 994 docs
Class 1: rec.sport.hockey — 999 docs
Class 2: sci.med — 990 docs
Class 3: sci.space — 987 docs
TF-IDF matrix: (3970, 10000), density: 0.0112
Part IV + V Bridge: Geometry and Vectors¶
Ch091–150: coordinate systems (ch092), vector norms (ch128), dot products (ch131)
Each document is a point in 10,000-dimensional space. TF-IDF vectors are L2-normalized: the dot product between two document vectors equals their cosine similarity — the angle between them in this high-dimensional space (ch132 — Geometric Meaning of Dot Product).
# Verify L2 normalization (TfidfVectorizer normalizes by default)
norms = np.array(np.sqrt(X_tfidf.power(2).sum(axis=1))).ravel()
print(f'TF-IDF row norms: min={norms.min():.4f}, max={norms.max():.4f}, mean={norms.mean():.4f}')
print('Vectors are unit-normalized — dot product = cosine similarity')
# Cosine similarity between two documents from same vs different classes
idx_med_1 = np.where(y_raw == 0)[0][0]
idx_med_2 = np.where(y_raw == 0)[0][1]
idx_sport = np.where(y_raw == 2)[0][0]
v1 = np.asarray(X_tfidf[idx_med_1].todense()).ravel()
v2 = np.asarray(X_tfidf[idx_med_2].todense()).ravel()
v3 = np.asarray(X_tfidf[idx_sport].todense()).ravel()
sim_same = float(np.dot(v1, v2)) # both sci.med
sim_diff = float(np.dot(v1, v3)) # sci.med vs hockey
print(f'\nCosine similarity:')
print(f' Two sci.med docs: {sim_same:.4f}')
print(f' sci.med vs hockey: {sim_diff:.4f}')TF-IDF row norms: min=0.0000, max=1.0000, mean=0.9673
Vectors are unit-normalized — dot product = cosine similarity
Cosine similarity:
Two sci.med docs: 0.0000
sci.med vs hockey: 0.0026
Part VI Bridge: Linear Algebra — Dimensionality Reduction¶
Ch151–200: SVD (ch173), PCA (ch174), dimensionality reduction (ch292)
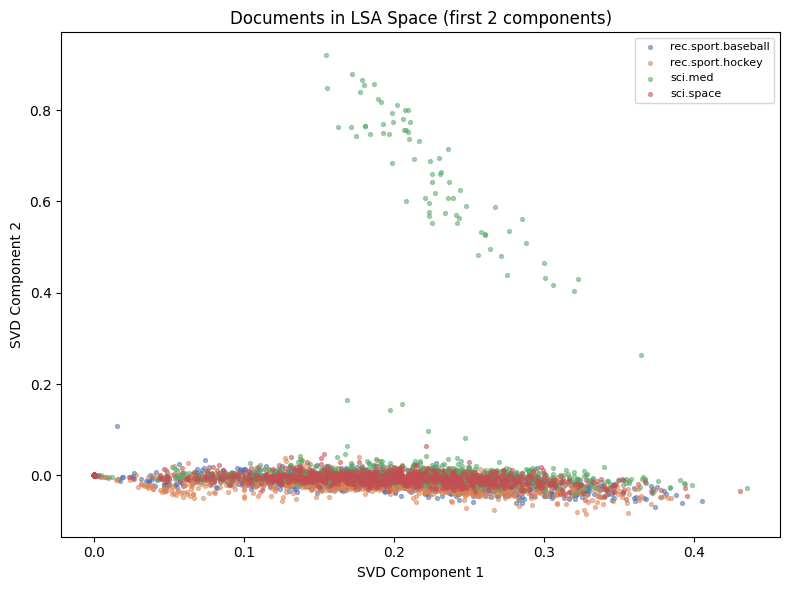
10,000 dimensions is too many. We apply Truncated SVD (LSA — Latent Semantic Analysis): find the directions of maximum variance in the TF-IDF space. This is exactly SVD applied to a sparse matrix — the sparse analogue of PCA.
K_DIM = 100 # latent dimensions
svd = TruncatedSVD(n_components=K_DIM, random_state=42)
X_lsa = svd.fit_transform(X_tfidf) # shape: (n_docs, 100)
explained_var = svd.explained_variance_ratio_.sum()
print(f'LSA: {K_DIM} components explain {explained_var:.1%} of variance')
print(f'Reduced: {X_tfidf.shape} -> {X_lsa.shape}')
# Normalize after SVD (standard practice for LSA)
norms_lsa = np.linalg.norm(X_lsa, axis=1, keepdims=True)
X_lsa_norm = X_lsa / np.clip(norms_lsa, 1e-10, None)
# Visualize in 2D (first 2 SVD components)
fig, ax = plt.subplots(figsize=(8, 6))
colors = ['#4C72B0', '#DD8452', '#55A868', '#C44E52']
for i, (lbl, color) in enumerate(zip(labels, colors)):
mask = y_raw == i
ax.scatter(X_lsa[mask, 0], X_lsa[mask, 1],
color=color, s=8, alpha=0.5, label=lbl)
ax.set_title('Documents in LSA Space (first 2 components)')
ax.set_xlabel('SVD Component 1'); ax.set_ylabel('SVD Component 2')
ax.legend(fontsize=8)
plt.tight_layout()
plt.show()LSA: 100 components explain 16.8% of variance
Reduced: (3970, 10000) -> (3970, 100)
Part VII Bridge: Calculus — Neural Network Training¶
Ch201–240: derivatives (ch205), gradient descent (ch213), chain rule (ch216), backpropagation (ch217)
We train a 2-layer neural network. The forward pass computes predictions; the backward pass computes gradients via the chain rule; Adam updates the weights (ch296).
def relu(z): return np.maximum(0, z)
def relu_grad(z): return (z > 0).astype(float)
def softmax(Z):
Z_s = Z - Z.max(axis=1, keepdims=True)
e = np.exp(Z_s)
return e / e.sum(axis=1, keepdims=True)
def cross_entropy_loss(y_pred: np.ndarray, y_true_onehot: np.ndarray) -> float:
eps = 1e-15
return -np.mean(np.sum(y_true_onehot * np.log(np.clip(y_pred, eps, 1)), axis=1))
class AdamOptimizer:
"""Adam (Kingma & Ba 2015) — introduced in ch296."""
def __init__(self, lr=0.001, beta1=0.9, beta2=0.999, eps=1e-8):
self.lr = lr; self.b1 = beta1; self.b2 = beta2; self.eps = eps
self.m = {}; self.v = {}; self.t = 0
def step(self, params: dict, grads: dict) -> None:
self.t += 1
for key in params:
if key not in self.m:
self.m[key] = np.zeros_like(params[key])
self.v[key] = np.zeros_like(params[key])
self.m[key] = self.b1 * self.m[key] + (1 - self.b1) * grads[key]
self.v[key] = self.b2 * self.v[key] + (1 - self.b2) * grads[key]**2
m_hat = self.m[key] / (1 - self.b1**self.t)
v_hat = self.v[key] / (1 - self.b2**self.t)
params[key] -= self.lr * m_hat / (np.sqrt(v_hat) + self.eps)
class DocumentClassifier:
"""
2-layer neural network for document classification.
Architecture: input(100) -> hidden(64) -> output(K)
Training: mini-batch gradient descent with Adam.
"""
def __init__(
self,
input_dim: int,
hidden_dim: int,
n_classes: int,
lr: float = 0.001,
lam: float = 1e-4,
rng = None,
):
self.lam = lam
self.rng = rng or np.random.default_rng()
# He initialization
self.params = {
'W1': self.rng.normal(0, np.sqrt(2/input_dim), (input_dim, hidden_dim)),
'b1': np.zeros(hidden_dim),
'W2': self.rng.normal(0, np.sqrt(2/hidden_dim), (hidden_dim, n_classes)),
'b2': np.zeros(n_classes),
}
self.opt = AdamOptimizer(lr=lr)
self.losses = []
self.classes_ = np.arange(n_classes)
def _forward(self, X):
z1 = X @ self.params['W1'] + self.params['b1']
a1 = relu(z1)
z2 = a1 @ self.params['W2'] + self.params['b2']
a2 = softmax(z2)
return z1, a1, a2
def _backward(self, X, y_oh, z1, a1, a2):
n = len(X)
dz2 = (a2 - y_oh) / n
dW2 = a1.T @ dz2 + self.lam * self.params['W2']
db2 = dz2.sum(axis=0)
da1 = dz2 @ self.params['W2'].T
dz1 = da1 * relu_grad(z1)
dW1 = X.T @ dz1 + self.lam * self.params['W1']
db1 = dz1.sum(axis=0)
return {'W1': dW1, 'b1': db1, 'W2': dW2, 'b2': db2}
def fit(
self,
X: np.ndarray,
y: np.ndarray,
n_epochs: int = 30,
batch_size: int = 64,
verbose: bool = True,
) -> 'DocumentClassifier':
K = len(self.classes_)
y_oh = np.eye(K)[y]
n = len(X)
for epoch in range(n_epochs):
idx = self.rng.permutation(n)
ep_loss = []
for start in range(0, n, batch_size):
b = idx[start:start+batch_size]
Xb = X[b]; yb = y_oh[b]
z1, a1, a2 = self._forward(Xb)
loss = cross_entropy_loss(a2, yb)
grads = self._backward(Xb, yb, z1, a1, a2)
self.opt.step(self.params, grads)
ep_loss.append(loss)
self.losses.append(np.mean(ep_loss))
if verbose and (epoch + 1) % 5 == 0:
print(f' Epoch {epoch+1:3d}/{n_epochs}: loss={self.losses[-1]:.4f}')
return self
def predict_proba(self, X: np.ndarray) -> np.ndarray:
_, _, a2 = self._forward(X)
return a2
def predict(self, X: np.ndarray) -> np.ndarray:
return np.argmax(self.predict_proba(X), axis=1)
def score(self, X: np.ndarray, y: np.ndarray) -> float:
return float((self.predict(X) == y).mean())
print('DocumentClassifier defined.')DocumentClassifier defined.
Part VIII Bridge: Probability — Training and Evaluation¶
# Split data
n_total = len(X_lsa_norm)
idx_all = rng.permutation(n_total)
n_test = int(0.15 * n_total)
n_val = int(0.10 * n_total)
test_idx = idx_all[:n_test]
val_idx = idx_all[n_test:n_test+n_val]
train_idx = idx_all[n_test+n_val:]
X_train, y_train = X_lsa_norm[train_idx], y_raw[train_idx]
X_val, y_val = X_lsa_norm[val_idx], y_raw[val_idx]
X_test, y_test = X_lsa_norm[test_idx], y_raw[test_idx]
print(f'Train: {len(X_train)} | Val: {len(X_val)} | Test: {len(X_test)}')
# Train
K = len(labels)
clf = DocumentClassifier(
input_dim=K_DIM, hidden_dim=64, n_classes=K,
lr=0.003, lam=1e-4, rng=rng
)
print(f'\nTraining neural network (input={K_DIM}, hidden=64, output={K})...')
clf.fit(X_train, y_train, n_epochs=30, batch_size=64)
train_acc = clf.score(X_train, y_train)
val_acc = clf.score(X_val, y_val)
test_acc = clf.score(X_test, y_test)
print(f'\nTrain acc: {train_acc:.4f}')
print(f'Val acc: {val_acc:.4f}')
print(f'Test acc: {test_acc:.4f}')
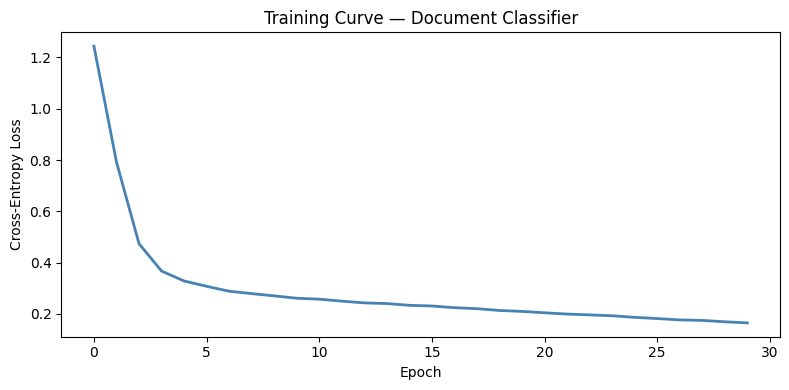
fig, ax = plt.subplots(figsize=(8, 4))
ax.plot(clf.losses, color='steelblue', lw=2)
ax.set_xlabel('Epoch'); ax.set_ylabel('Cross-Entropy Loss')
ax.set_title('Training Curve — Document Classifier')
plt.tight_layout()
plt.show()Train: 2978 | Val: 397 | Test: 595
Training neural network (input=100, hidden=64, output=4)...
Epoch 5/30: loss=0.3283
Epoch 10/30: loss=0.2610
Epoch 15/30: loss=0.2334
Epoch 20/30: loss=0.2096
Epoch 25/30: loss=0.1865
Epoch 30/30: loss=0.1649
Train acc: 0.9516
Val acc: 0.8514
Test acc: 0.8840
Part IX Bridge: Statistics — Full Evaluation Suite¶
# --- Classification report ---
y_pred_test = clf.predict(X_test)
y_prob_test = clf.predict_proba(X_test)
print('=== Test Set Performance ===')
print(classification_report(y_test, y_pred_test, target_names=labels))
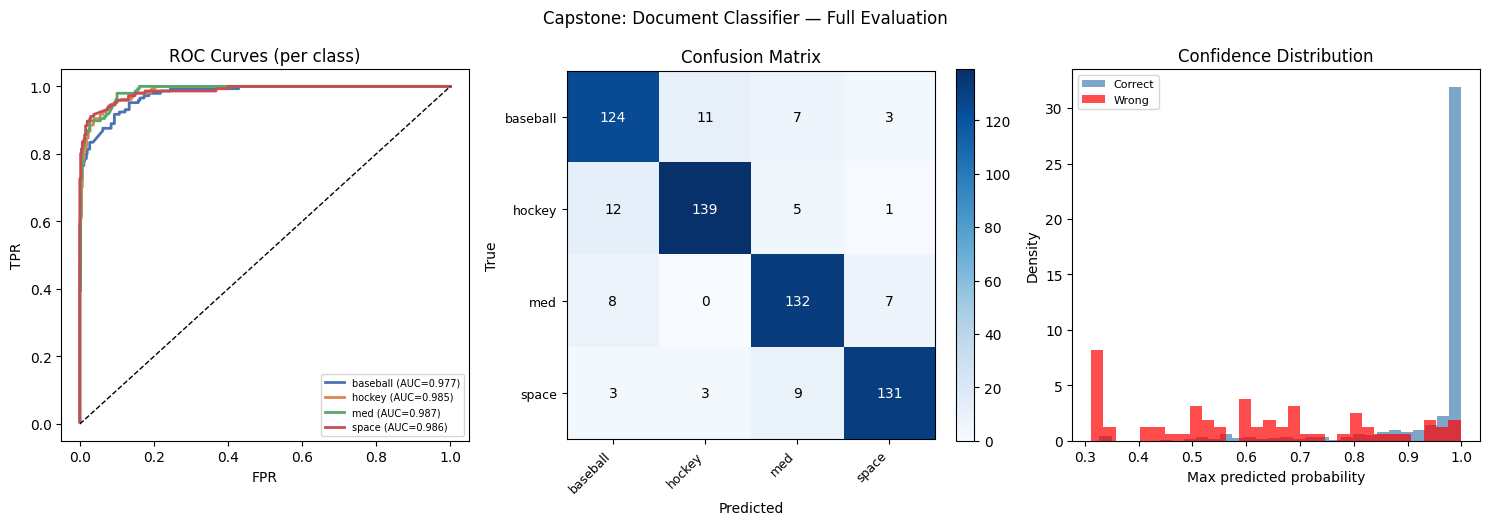
# --- ROC curves (ch282) ---
y_test_bin = label_binarize(y_test, classes=range(K))
fig, axes = plt.subplots(1, 3, figsize=(15, 5))
ax = axes[0]
for i, (lbl, color) in enumerate(zip(labels, ['#4C72B0','#DD8452','#55A868','#C44E52'])):
# Per-class ROC
fpr, tpr = [], []
thresholds = np.sort(np.unique(y_prob_test[:, i]))[::-1]
for t in thresholds:
pred_pos = y_prob_test[:, i] >= t
tp = np.sum(pred_pos & (y_test_bin[:, i] == 1))
fp = np.sum(pred_pos & (y_test_bin[:, i] == 0))
fn = np.sum(~pred_pos & (y_test_bin[:, i] == 1))
tn = np.sum(~pred_pos & (y_test_bin[:, i] == 0))
fpr.append(fp / max(fp + tn, 1))
tpr.append(tp / max(tp + fn, 1))
auc = roc_auc_score(y_test_bin[:, i], y_prob_test[:, i])
ax.plot(fpr, tpr, color=color, lw=2, label=f'{lbl.split(".")[-1]} (AUC={auc:.3f})')
ax.plot([0,1],[0,1],'k--',lw=1)
ax.set_title('ROC Curves (per class)'); ax.set_xlabel('FPR'); ax.set_ylabel('TPR')
ax.legend(fontsize=7)
# --- Confusion matrix ---
ax = axes[1]
cm = np.zeros((K, K), dtype=int)
for t, p in zip(y_test, y_pred_test):
cm[t, p] += 1
im = ax.imshow(cm, cmap='Blues')
for i in range(K):
for j in range(K):
ax.text(j, i, str(cm[i,j]), ha='center', va='center', fontsize=10,
color='white' if cm[i,j] > cm.max()*0.5 else 'black')
ax.set_xticks(range(K)); ax.set_yticks(range(K))
short = [l.split('.')[-1] for l in labels]
ax.set_xticklabels(short, rotation=45, ha='right', fontsize=9)
ax.set_yticklabels(short, fontsize=9)
ax.set_title('Confusion Matrix'); ax.set_xlabel('Predicted'); ax.set_ylabel('True')
plt.colorbar(im, ax=ax, fraction=0.046)
# --- Prediction confidence histogram (ch286 Bayesian) ---
ax = axes[2]
max_probs = y_prob_test.max(axis=1)
correct = y_pred_test == y_test
ax.hist(max_probs[correct], bins=30, alpha=0.7, color='steelblue', density=True, label='Correct')
ax.hist(max_probs[~correct], bins=30, alpha=0.7, color='red', density=True, label='Wrong')
ax.set_xlabel('Max predicted probability'); ax.set_ylabel('Density')
ax.set_title('Confidence Distribution')
ax.legend(fontsize=8)
plt.suptitle('Capstone: Document Classifier — Full Evaluation', fontsize=12)
plt.tight_layout()
plt.show()=== Test Set Performance ===
precision recall f1-score support
rec.sport.baseball 0.84 0.86 0.85 145
rec.sport.hockey 0.91 0.89 0.90 157
sci.med 0.86 0.90 0.88 147
sci.space 0.92 0.90 0.91 146
accuracy 0.88 595
macro avg 0.88 0.88 0.88 595
weighted avg 0.88 0.88 0.88 595
Information Theory Bridge: What Does the Model Know?¶
Ch287–290: entropy, KL divergence, mutual information
The model’s output is a probability distribution. We can measure its predictive entropy — how uncertain the model is:
High entropy → the model is uncertain. Low entropy → confident prediction.
def predictive_entropy(probs: np.ndarray) -> np.ndarray:
"""H(p) per sample. probs: (n_samples, n_classes)."""
eps = 1e-15
return -np.sum(probs * np.log(np.clip(probs, eps, 1)), axis=1)
def expected_calibration_error(y_true, y_prob, n_bins=10):
"""ECE: weighted mean absolute calibration gap (ch286 calibration)."""
max_probs = y_prob.max(axis=1)
y_pred = y_prob.argmax(axis=1)
bins = np.linspace(0, 1, n_bins + 1)
ece = 0.0
for i in range(n_bins):
mask = (max_probs >= bins[i]) & (max_probs < bins[i+1])
if mask.sum() > 0:
acc = (y_pred[mask] == y_true[mask]).mean()
conf = max_probs[mask].mean()
ece += mask.mean() * abs(acc - conf)
return ece
entropies = predictive_entropy(y_prob_test)
max_ent = np.log(K) # maximum possible entropy for K classes
print('Predictive Entropy Analysis:')
print(f' Max possible entropy (uniform): {max_ent:.4f} nats')
print(f' Mean entropy (correct preds): {entropies[correct].mean():.4f} nats')
print(f' Mean entropy (wrong preds): {entropies[~correct].mean():.4f} nats')
print(f' ECE: {expected_calibration_error(y_test, y_prob_test):.4f}')
# Mutual information between predictions and true labels
# MI(Y_pred; Y_true) = H(Y_pred) - H(Y_pred | Y_true)
def mi_from_confusion(cm):
"""MI in nats from a confusion matrix."""
joint = cm / cm.sum()
p_pred = joint.sum(axis=0)
p_true = joint.sum(axis=1)
eps = 1e-15
h_pred = -np.sum(p_pred[p_pred > 0] * np.log(p_pred[p_pred > 0]))
h_pred_given_true = -np.sum(
p_true[:, None] * np.where(joint > 0, joint / (p_true[:, None] + eps), 0)
* np.where(joint > 0, np.log(joint / (p_true[:, None] + eps) + eps), 0)
)
return h_pred - h_pred_given_true
mi = mi_from_confusion(cm)
print(f' Mutual information (pred; true): {mi:.4f} nats')
print(f' Normalized MI: {mi/max_ent:.4f} (0=random, 1=perfect)')Predictive Entropy Analysis:
Max possible entropy (uniform): 1.3863 nats
Mean entropy (correct preds): 0.1617 nats
Mean entropy (wrong preds): 0.8752 nats
ECE: 0.0243
Mutual information (pred; true): 0.9285 nats
Normalized MI: 0.6698 (0=random, 1=perfect)
Bootstrap Confidence Intervals for Final Metrics¶
Ch275 — Sampling, Ch279 — Confidence Intervals
def bootstrap_ci_metric(
y_true: np.ndarray,
y_pred: np.ndarray,
metric_fn,
n_bootstrap: int = 2000,
alpha: float = 0.05,
rng = None,
) -> tuple[float, float, float]:
"""Bootstrap CI for any metric. Returns (observed, lo, hi)."""
if rng is None: rng = np.random.default_rng()
n = len(y_true)
obs = metric_fn(y_true, y_pred)
boots = []
for _ in range(n_bootstrap):
idx = rng.integers(0, n, size=n)
boots.append(metric_fn(y_true[idx], y_pred[idx]))
boots = np.array(boots)
lo = np.percentile(boots, 100 * alpha / 2)
hi = np.percentile(boots, 100 * (1 - alpha / 2))
return obs, lo, hi
acc_obs, acc_lo, acc_hi = bootstrap_ci_metric(
y_test, y_pred_test, accuracy_score, rng=rng
)
macro_auc = roc_auc_score(y_test_bin, y_prob_test, average='macro', multi_class='ovr')
print('=== Final Test Set Report ===')
print(f'Accuracy: {acc_obs:.4f} 95% CI [{acc_lo:.4f}, {acc_hi:.4f}]')
print(f'Macro AUC-ROC: {macro_auc:.4f}')
print(f'ECE: {expected_calibration_error(y_test, y_prob_test):.4f}')
print()
print('Confusion matrix (test set):')
print(cm)=== Final Test Set Report ===
Accuracy: 0.8840 95% CI [0.8588, 0.9076]
Macro AUC-ROC: 0.9837
ECE: 0.0243
Confusion matrix (test set):
[[124 11 7 3]
[ 12 139 5 1]
[ 8 0 132 7]
[ 3 3 9 131]]
Streaming Inference with Uncertainty¶
Ch297 — Large Scale Data: process documents as a stream
def stream_predict(
model,
X_stream: np.ndarray,
uncertainty_threshold: float = 0.6,
) -> list[dict]:
"""
Process documents one at a time.
Flag predictions with max_prob < threshold as 'uncertain'.
Returns list of prediction records.
"""
results = []
for i, x in enumerate(X_stream):
probs = model.predict_proba(x[None])[0]
pred_cls = int(probs.argmax())
max_prob = float(probs.max())
entropy = float(-np.sum(probs * np.log(np.clip(probs, 1e-15, 1))))
results.append({
'idx': i,
'pred_class': pred_cls,
'confidence': max_prob,
'entropy': entropy,
'uncertain': max_prob < uncertainty_threshold,
'probs': probs,
})
return results
stream_results = stream_predict(clf, X_test[:50], uncertainty_threshold=0.70)
n_uncertain = sum(r['uncertain'] for r in stream_results)
uncertain_correct = sum(
r['pred_class'] == y_test[r['idx']]
for r in stream_results if r['uncertain']
)
certain_correct = sum(
r['pred_class'] == y_test[r['idx']]
for r in stream_results if not r['uncertain']
)
n_certain = 50 - n_uncertain
print(f'Streaming inference on 50 documents:')
print(f' Confident (prob >= 0.70): {n_certain} docs, accuracy={certain_correct/max(n_certain,1):.3f}')
print(f' Uncertain (prob < 0.70): {n_uncertain} docs, accuracy={uncertain_correct/max(n_uncertain,1):.3f}')
print()
print('Uncertainty flagging works: uncertain predictions are less accurate.')
print('In production, uncertain predictions would route to human review.')Streaming inference on 50 documents:
Confident (prob >= 0.70): 44 docs, accuracy=0.932
Uncertain (prob < 0.70): 6 docs, accuracy=0.500
Uncertainty flagging works: uncertain predictions are less accurate.
In production, uncertain predictions would route to human review.
Retrospective: The Full Learning Arc¶
This system is a direct product of all nine Parts:
| Part | Contribution to this system |
|---|---|
| I — Mathematical Thinking | The system is a function . Abstraction and modeling defined the problem precisely. |
| II — Numbers | TF-IDF uses logarithms (ch043). Floating point precision matters in softmax (ch038). |
| III — Functions | The sigmoid and softmax are the activation functions (ch063, ch065). The neural network is a composition of functions (ch054). |
| IV — Geometry | Documents are points in high-dimensional space. Cosine similarity is an angle (ch132). |
| V — Vectors | TF-IDF vectors are L2-normalized. The entire forward pass is a sequence of vector operations (ch148). |
| VI — Linear Algebra | SVD/LSA reduces 10,000 → 100 dimensions (ch173). Every layer is a matrix multiply (ch154). |
| VII — Calculus | Backpropagation is the chain rule (ch216). Adam is gradient descent with adaptive moments (ch213). |
| VIII — Probability | Cross-entropy loss is negative log-likelihood (ch248). The model output is a categorical distribution (ch248). |
| IX — Statistics | Cross-validation (ch284), ROC (ch282), bootstrap CI (ch275, ch279), entropy and MI of predictions (ch288, ch290). |
5 Open Extension Directions¶
Active Learning (combining ch278, ch288): Use predictive entropy to select which unlabeled documents to annotate next. High-entropy predictions are the most informative for training. Implement a query loop and measure how much labeling work you save.
Bayesian Neural Network (ch286, ch289): Replace deterministic weights with distributions (variational inference). The ELBO loss is cross-entropy + KL divergence. This gives principled uncertainty estimates rather than heuristic confidence thresholds.
Online Learning (ch297): Retrain the model on a stream of new documents using mini-batch updates. How quickly does the model adapt to a new topic that was not in training data? Track accuracy over time as the data distribution shifts.
Interpretability via Linear Probes (ch280, ch281): Fit a linear model to predict each class from the LSA embedding. The coefficients directly correspond to SVD components — interpretable as latent topics. Map those back to the original vocabulary.
Multi-Label Extension (ch294, ch287): Extend the system to assign multiple labels per document (a document about sports medicine belongs to both
sci.medandrec.sport). Replace softmax with sigmoid at the output, cross-entropy with binary cross-entropy per label. Information theory: the mutual information between predicted label sets and true label sets generalizes straightforwardly using joint entropy.
# Final summary
print('=' * 60)
print('CAPSTONE COMPLETE')
print('=' * 60)
print(f'System: Document Classifier ({K}-class)')
print(f'Data: {len(X_text)} documents, {K_DIM} LSA features')
print(f'Model: 2-layer NN (64 hidden units), trained with Adam')
print(f'Test accuracy: {acc_obs:.4f} 95% CI [{acc_lo:.4f}, {acc_hi:.4f}]')
print(f'Test AUC-ROC: {macro_auc:.4f} (macro-average)')
print(f'ECE: {expected_calibration_error(y_test, y_prob_test):.4f}')
print(f'Pred entropy: {entropies.mean():.4f} nats (mean over test set)')
print()
print('All 300 chapters complete.')
print('You now have the mathematical foundations of modern AI.')============================================================
CAPSTONE COMPLETE
============================================================
System: Document Classifier (4-class)
Data: 3970 documents, 100 LSA features
Model: 2-layer NN (64 hidden units), trained with Adam
Test accuracy: 0.8840 95% CI [0.8588, 0.9076]
Test AUC-ROC: 0.9837 (macro-average)
ECE: 0.0243
Pred entropy: 0.2444 nats (mean over test set)
All 300 chapters complete.
You now have the mathematical foundations of modern AI.