1. What a loss function is¶
A loss function measures the cost of predicting when the true label is . The training loss is its average over the dataset:
Loss function design is not arbitrary. A good loss must:
Be differentiable almost everywhere (gradient descent requires ).
Have a minimum at the correct prediction (otherwise the gradient points in the wrong direction).
Produce informative gradients even for nearly-correct predictions.
Be statistically principled — ideally derived from maximum likelihood estimation.
2. Mean Squared Error (regression)¶
Gradient: . Linear in the residual — large errors produce large gradients, small errors produce small gradients.
Statistical derivation: MSE is the negative log-likelihood under a Gaussian noise model: . Minimising MSE = maximising likelihood.
(Gaussian distribution: ch253. Log-likelihood: ch293.)
3. Binary Cross-Entropy (binary classification)¶
where is the predicted probability and .
Why not MSE for classification? If the output is a probability from a sigmoid, MSE gradient nearly vanishes when is confident but wrong (sigmoid saturated). BCE gradient: — clean, non-vanishing.
(Entropy and KL divergence: ch294–ch295. Sigmoid: ch064.)
4. Categorical Cross-Entropy (multiclass)¶
where and is one-hot encoded.
Softmax:
Combined gradient of CE + softmax: . This elegant result is why CE+softmax is the universal choice for classification.
import numpy as np
import matplotlib.pyplot as plt
def mse(y_hat: np.ndarray, y: np.ndarray) -> float:
return float(np.mean((y_hat - y) ** 2))
def bce(p_hat: np.ndarray, y: np.ndarray, eps: float = 1e-12) -> float:
"""Binary cross-entropy. Clips to avoid log(0)."""
p = np.clip(p_hat, eps, 1 - eps)
return float(-np.mean(y * np.log(p) + (1 - y) * np.log(1 - p)))
def softmax(z: np.ndarray) -> np.ndarray:
"""Numerically stable softmax using log-sum-exp trick."""
z_shifted = z - z.max(axis=0, keepdims=True)
exp_z = np.exp(z_shifted)
return exp_z / exp_z.sum(axis=0, keepdims=True)
def cross_entropy(logits: np.ndarray, y_onehot: np.ndarray, eps: float = 1e-12) -> float:
"""Categorical cross-entropy from logits."""
p = softmax(logits)
return float(-np.mean(np.sum(y_onehot * np.log(p + eps), axis=0)))
# --- Visualise BCE vs MSE gradient behaviour ---
p = np.linspace(0.01, 0.99, 300)
# True label y=1
bce_loss = -(1 * np.log(p) + 0 * np.log(1 - p))
mse_loss = (1 - p) ** 2
# Gradients w.r.t. z (pre-sigmoid), using chain rule
# d(sigmoid)/dz = p*(1-p)
dbce_dz = p - 1 # p - y (when y=1)
dmse_dz = 2 * (p - 1) * p * (1 - p) # MSE grad × sigmoid derivative
fig, axes = plt.subplots(2, 2, figsize=(12, 8))
for ax, loss, name, color in [
(axes[0, 0], bce_loss, 'Binary Cross-Entropy (y=1)', '#e74c3c'),
(axes[0, 1], mse_loss, 'MSE (y=1)', '#3498db'),
]:
ax.plot(p, loss, color=color, lw=2)
ax.set_xlabel('Predicted probability $\\hat{p}$')
ax.set_ylabel('Loss')
ax.set_title(name)
ax.axvline(1.0, color='green', linestyle='--', alpha=0.5, label='Correct (p=1)')
ax.legend()
for ax, grad, name, color in [
(axes[1, 0], dbce_dz, 'BCE gradient w.r.t. z', '#e74c3c'),
(axes[1, 1], dmse_dz, 'MSE gradient w.r.t. z', '#3498db'),
]:
ax.plot(p, grad, color=color, lw=2)
ax.axhline(0, color='black', alpha=0.3)
ax.set_xlabel('Predicted probability $\\hat{p}$')
ax.set_ylabel('Gradient')
ax.set_title(name)
ax.fill_between(p, grad, 0, alpha=0.15, color=color)
plt.suptitle('BCE has informative gradients even at extreme predictions;\n'
'MSE gradient vanishes at saturation.', fontsize=11)
plt.tight_layout()
plt.savefig('ch305_loss_gradients.png', dpi=120)
plt.show()
# Demonstration: gradient magnitude at nearly-wrong confident prediction
p_wrong = 0.99 # predicted 0.99 but true label is 0
bce_grad = p_wrong - 0
mse_grad = 2 * (p_wrong - 0) * p_wrong * (1 - p_wrong)
print(f"Confident wrong prediction (p={p_wrong}, y=0):")
print(f" BCE gradient |dL/dz| = {abs(bce_grad):.4f} ← still strong")
print(f" MSE gradient |dL/dz| = {abs(mse_grad):.4f} ← nearly zero!")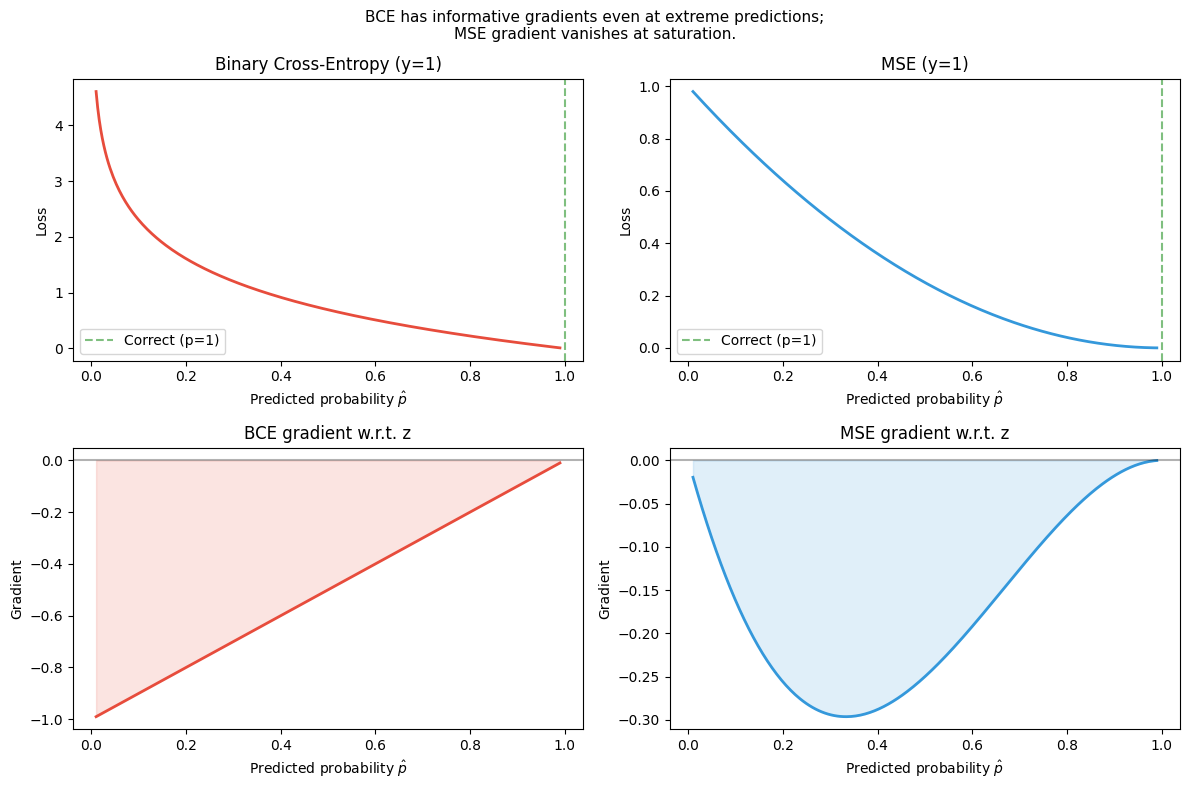
Confident wrong prediction (p=0.99, y=0):
BCE gradient |dL/dz| = 0.9900 ← still strong
MSE gradient |dL/dz| = 0.0196 ← nearly zero!
# --- Softmax behaviour ---
logits = np.array([[2.0, 1.0, 0.1, -1.0],
[5.0, 1.0, 0.1, -1.0],
[50.0, 1.0, 0.1, -1.0]])
print("Softmax concentrates probability mass as the winning logit grows:")
for z in logits:
p = softmax(z[:, None]).ravel()
print(f" logits={z} → p={np.round(p, 3)}")
# Numerically UNSTABLE version for comparison
def softmax_unstable(z):
exp_z = np.exp(z)
return exp_z / exp_z.sum()
large_z = np.array([1000.0, 999.0, 998.0])
print(f"\nUnstable softmax on large logits: {softmax_unstable.__name__}")
try:
result = softmax_unstable(large_z)
print(f" Result: {result} (overflow produces NaN/inf if not handled)")
except Exception as e:
print(f" Exception: {e}")
stable_result = softmax(large_z[:, None]).ravel()
print(f"Stable softmax (log-sum-exp): {stable_result}")Softmax concentrates probability mass as the winning logit grows:
logits=[ 2. 1. 0.1 -1. ] → p=[0.638 0.235 0.095 0.032]
logits=[ 5. 1. 0.1 -1. ] → p=[0.973 0.018 0.007 0.002]
logits=[50. 1. 0.1 -1. ] → p=[1. 0. 0. 0.]
Unstable softmax on large logits: softmax_unstable
Result: [nan nan nan] (overflow produces NaN/inf if not handled)
Stable softmax (log-sum-exp): [0.66524096 0.24472847 0.09003057]
C:\Users\user\AppData\Local\Temp\ipykernel_25912\3083791601.py:13: RuntimeWarning: overflow encountered in exp
exp_z = np.exp(z)
C:\Users\user\AppData\Local\Temp\ipykernel_25912\3083791601.py:14: RuntimeWarning: invalid value encountered in divide
return exp_z / exp_z.sum()
5. Huber loss (robust regression)¶
MSE is sensitive to outliers because grows quadratically. Huber loss interpolates between MSE (for small errors) and MAE (for large errors):
Differentiable everywhere; gradient bounded by . Used in reinforcement learning (ch329) where reward estimates have heavy-tailed noise.
6. Choosing the loss function¶
| Task | Output activation | Loss |
|---|---|---|
| Regression | Linear | MSE |
| Robust regression | Linear | Huber |
| Binary classification | Sigmoid | BCE |
| Multiclass (mutually exclusive) | Softmax | Categorical CE |
| Multilabel classification | Sigmoid (per class) | BCE (per class) |
| Generative models | Varies | KL divergence, ELBO |
7. Summary¶
Loss functions must be differentiable and have informative gradients.
MSE = negative log-likelihood under Gaussian noise. BCE = negative log-likelihood for Bernoulli.
The BCE+sigmoid and CE+softmax gradients simplify to — elegant and non-vanishing.
Loss choice encodes assumptions about the noise distribution of the labels.
8. Forward and backward references¶
Used here: entropy (ch294), KL divergence (ch295), Gaussian distribution (ch253), log-likelihood (ch293), sigmoid (ch064), derivatives (ch205–ch206).
This will reappear in ch306 — Backpropagation from Scratch, where the loss gradient is the starting point of the backward pass, and in ch325 — VAE, where the ELBO loss combines reconstruction CE and KL divergence.