1. The problem with vanilla RNNs¶
As shown in ch317, gradients vanish over long sequences because involves repeated multiplication by . The network cannot learn dependencies spanning many time steps.
Long Short-Term Memory (Hochreiter & Schmidhuber, 1997) introduces a cell state that flows through time with only additive interactions — no repeated matrix multiplication — allowing gradients to propagate over hundreds of steps.
2. LSTM equations¶
The cell state is modified only by addition — the constant error carousel that allows gradients to flow without decay over long sequences.
(Sigmoid: ch309. Element-wise operations: ch123.)
import numpy as np
import matplotlib.pyplot as plt
def sigmoid(z): return 1/(1+np.exp(-np.clip(z,-500,500)))
class LSTMCell:
"""Single LSTM cell (unrolled for clarity)."""
def __init__(self, input_dim: int, hidden_dim: int, seed: int = 0):
rng = np.random.default_rng(seed)
H = hidden_dim; concat = input_dim + hidden_dim
s = 1.0 / np.sqrt(concat)
# All 4 gate weight matrices packed into one for efficiency
self.W = rng.normal(0, s, (4 * H, concat))
self.b = np.zeros(4 * H)
# Initialise forget gate bias to 1 for better gradient flow
self.b[H:2*H] = 1.0
self.H = H
def forward(self, x: np.ndarray, h_prev: np.ndarray,
c_prev: np.ndarray) -> tuple:
H = self.H
xh = np.concatenate([x, h_prev])
gates = self.W @ xh + self.b
i = sigmoid(gates[:H]) # input gate
f = sigmoid(gates[H:2*H]) # forget gate
o = sigmoid(gates[2*H:3*H]) # output gate
g = np.tanh(gates[3*H:]) # candidate
c = f * c_prev + i * g # cell state update (additive!)
h = o * np.tanh(c) # hidden state
return h, c, (xh, i, f, o, g, c_prev, c)
class GRUCell:
"""Gated Recurrent Unit — simpler than LSTM, often comparable."""
def __init__(self, input_dim: int, hidden_dim: int, seed: int = 0):
rng = np.random.default_rng(seed)
H = hidden_dim; concat = input_dim + hidden_dim
s = 1.0 / np.sqrt(concat)
self.W_z = rng.normal(0, s, (H, concat)) # update gate
self.W_r = rng.normal(0, s, (H, concat)) # reset gate
self.W_h = rng.normal(0, s, (H, concat)) # candidate
self.b_z = np.zeros(H); self.b_r = np.zeros(H); self.b_h = np.zeros(H)
self.H = H
def forward(self, x: np.ndarray, h_prev: np.ndarray) -> np.ndarray:
xh = np.concatenate([x, h_prev])
z = sigmoid(self.W_z @ xh + self.b_z) # update gate
r = sigmoid(self.W_r @ xh + self.b_r) # reset gate
xh_r = np.concatenate([x, r * h_prev])
h_cand = np.tanh(self.W_h @ xh_r + self.b_h) # candidate
h = (1 - z) * h_prev + z * h_cand # interpolate old/new
return h
# ── Compare gradient flow: RNN vs LSTM ──
rng = np.random.default_rng(0)
T = 100; D = 4; H = 32
lstm = LSTMCell(D, H, seed=0)
x_seq = rng.normal(0, 1, (T, D))
# Track gate activations over time
h = np.zeros(H); c = np.zeros(H)
gate_log = {'i': [], 'f': [], 'o': [], 'g_norm': []}
for x in x_seq:
h, c, (_, i, f, o, g, _, _) = lstm.forward(x, h, c)
gate_log['i'].append(i.mean()); gate_log['f'].append(f.mean())
gate_log['o'].append(o.mean()); gate_log['g_norm'].append(np.linalg.norm(c))
fig, axes = plt.subplots(2, 2, figsize=(12, 7))
for ax, key, color, label in zip(axes.ravel(),
['i','f','o','g_norm'],
['#3498db','#e74c3c','#2ecc71','#9b59b6'],
['Input gate (mean)', 'Forget gate (mean)', 'Output gate (mean)', 'Cell state norm']):
ax.plot(gate_log[key], color=color, lw=1.5)
ax.set_title(label); ax.set_xlabel('Time step')
plt.suptitle('LSTM gate activations over 100 time steps', fontsize=11)
plt.tight_layout()
plt.savefig('ch318_lstm_gates.png', dpi=120)
plt.show()
# Parameter count comparison
print("Parameter comparison (D=4, H=32):")
lstm_params = 4 * H * (D + H) + 4 * H
gru_params = 3 * H * (D + H) + 3 * H
rnn_params = H * (D + H) + H
print(f" Vanilla RNN: {rnn_params:,}")
print(f" GRU: {gru_params:,}")
print(f" LSTM: {lstm_params:,}")
print(f" GRU/LSTM ratio: {gru_params/lstm_params:.2f} (GRU is 75% the size)")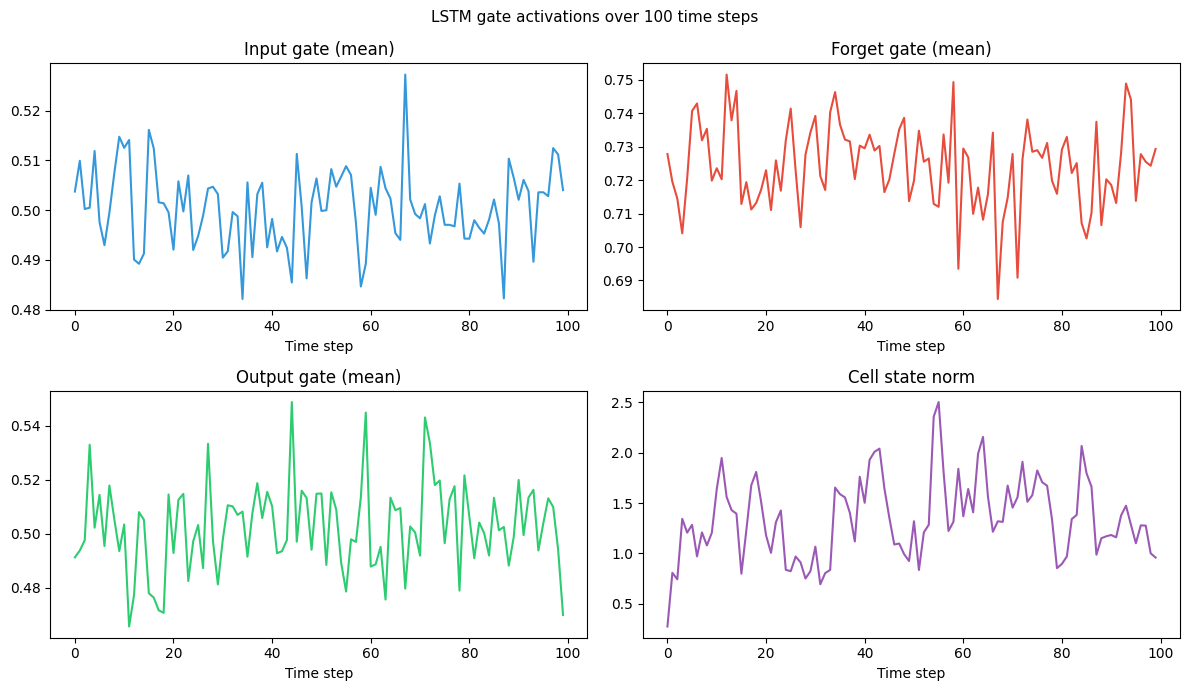
Parameter comparison (D=4, H=32):
Vanilla RNN: 1,184
GRU: 3,552
LSTM: 4,736
GRU/LSTM ratio: 0.75 (GRU is 75% the size)
3. Why the cell state solves vanishing gradients¶
The cell update rule is purely additive. The gradient of the loss with respect to an old cell state is:
This is a product of forget gate values — all in . But crucially, the forget gate can learn to set for dimensions that should be remembered, keeping that gradient close to 1 over any number of steps.
4. Summary¶
LSTM: input, forget, output gates + cell state. Additive cell update avoids vanishing gradients.
Forget gate bias initialised to 1: encourages remembering by default at training start.
GRU: update + reset gates, no separate cell state. Fewer parameters; similar performance.
Both are standard seq-to-seq building blocks, now largely superseded by Transformers for NLP.
5. Forward and backward references¶
Used here: sigmoid (ch309), tanh (ch309), element-wise products (ch123), vanishing gradients (ch317).
This will reappear in ch319 — Sequence-to-Sequence Models, where stacked LSTMs form encoder-decoder architectures, and in ch336 — Project: Character-Level LM.