Prerequisites: ch139 (Basis and Dimension), ch138 (Subspaces), ch127 (Linear Combination)
You will learn:
What the span of a set of vectors is and why it is always a subspace
How to determine whether a vector is in the span of a given set
How adding or removing vectors changes the span
The relationship between span, basis, and dimension
How span appears in practice in systems of equations and data science
Environment: Python 3.x, numpy, matplotlib
1. Concept¶
The span of a set of vectors S = {v₁, v₂, ..., vₖ} is the set of all linear combinations of those vectors:
span(S) = {c₁v₁ + c₂v₂ + ... + cₖvₖ : c₁, c₂, ..., cₖ ∈ ℝ}Span answers the question: what territory can I reach if I’m allowed to walk in the directions given by the vectors in S, scaling and combining freely?
Key facts:
The span of any set of vectors is always a subspace (verified in ch138).
The span of the empty set is {0} (the trivial subspace).
A set of vectors spans a space V if span(S) = V.
A basis is exactly a spanning set that is also linearly independent — the minimal spanning set.
Common misconceptions:
“Adding more vectors always increases the span.” False — adding a vector already in the span leaves it unchanged. A dependent vector contributes nothing new.
“The span of n vectors in ℝⁿ is ℝⁿ.” False — if the vectors are linearly dependent, the span is a strict subspace of ℝⁿ.
2. Intuition & Mental Models¶
Geometric model: Think of span as reachability. Stand at the origin. Each vector in S is a direction you can travel. You can go 3 steps in direction v₁, then -2 steps in direction v₂, then 0.5 steps in v₃ — any combination, any scale. The set of all positions you can reach is the span.
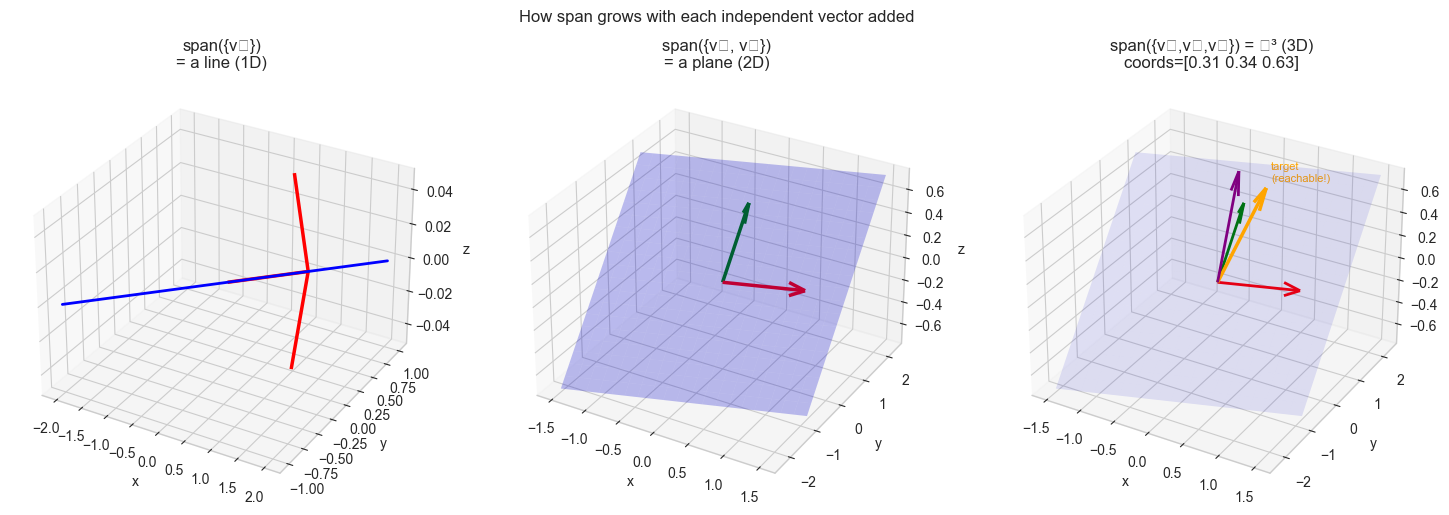
One non-zero vector spans a line through the origin.
Two non-parallel vectors in ℝ³ span a plane through the origin.
Three linearly independent vectors in ℝ³ span all of ℝ³.
Computational model: Span is the image of a matrix. If A is the matrix whose columns are v₁, ..., vₖ, then:
span({v₁, ..., vₖ}) = {Ax : x ∈ ℝᵏ} = column space C(A)The question “is b in the span?” becomes “does Ax = b have a solution?” — a linear system, which we know how to solve. (This will be formalized in ch160.)
Recall from ch127 (Linear Combination): every element of the span is a linear combination. The span is the closure of S under linear combination — you cannot leave it by combining vectors already in it.
3. Visualization¶
# --- Visualization: Span of 1, 2, and 3 vectors in R^3 ---
# Shows how span grows from a line -> plane -> all of R^3
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
plt.style.use('seaborn-v0_8-whitegrid')
fig = plt.figure(figsize=(15, 5))
v1 = np.array([1.0, 0.5, 0.0])
v2 = np.array([0.0, 1.0, 0.5])
v3 = np.array([0.3, 0.0, 1.0])
# --- Panel 1: span({v1}) — a line ---
ax1 = fig.add_subplot(131, projection='3d')
t = np.linspace(-2, 2, 100)
line = np.outer(t, v1)
ax1.plot(line[:, 0], line[:, 1], line[:, 2], 'b-', lw=2)
ax1.quiver(0, 0, 0, *v1, color='red', lw=2.5, arrow_length_ratio=0.2)
ax1.set_title('span({v₁})\n= a line (1D)')
ax1.set_xlabel('x'); ax1.set_ylabel('y'); ax1.set_zlabel('z')
# --- Panel 2: span({v1, v2}) — a plane ---
ax2 = fig.add_subplot(132, projection='3d')
s_vals = np.linspace(-1.5, 1.5, 20)
t_vals = np.linspace(-1.5, 1.5, 20)
S, T = np.meshgrid(s_vals, t_vals)
X = S * v1[0] + T * v2[0]
Y = S * v1[1] + T * v2[1]
Z = S * v1[2] + T * v2[2]
ax2.plot_surface(X, Y, Z, alpha=0.25, color='blue')
ax2.quiver(0,0,0, *v1, color='red', lw=2.5, arrow_length_ratio=0.2)
ax2.quiver(0,0,0, *v2, color='green', lw=2.5, arrow_length_ratio=0.2)
ax2.set_title('span({v₁, v₂})\n= a plane (2D)')
ax2.set_xlabel('x'); ax2.set_ylabel('y'); ax2.set_zlabel('z')
# --- Panel 3: span({v1, v2, v3}) — all of R^3 (if independent) ---
ax3 = fig.add_subplot(133, projection='3d')
# Show that we can reach a point off the v1-v2 plane
B = np.column_stack([v1, v2, v3])
target = np.array([0.5, 0.5, 0.8])
c = np.linalg.solve(B, target)
ax3.plot_surface(X, Y, Z, alpha=0.1, color='blue') # v1-v2 plane for reference
ax3.quiver(0,0,0,*v1,color='red',lw=2,arrow_length_ratio=0.2)
ax3.quiver(0,0,0,*v2,color='green',lw=2,arrow_length_ratio=0.2)
ax3.quiver(0,0,0,*v3,color='purple',lw=2,arrow_length_ratio=0.2)
ax3.quiver(0,0,0,*target,color='orange',lw=2.5,arrow_length_ratio=0.2)
ax3.text(*target+0.05, f'target\n(reachable!)', fontsize=8, color='orange')
ax3.set_title(f'span({{v₁,v₂,v₃}}) = ℝ³ (3D)\ncoords={c.round(2)}')
ax3.set_xlabel('x'); ax3.set_ylabel('y'); ax3.set_zlabel('z')
plt.suptitle('How span grows with each independent vector added', fontsize=12, y=1.01)
plt.tight_layout()
plt.show()C:\Users\user\AppData\Local\Temp\ipykernel_13356\850510082.py:55: UserWarning: Glyph 8321 (\N{SUBSCRIPT ONE}) missing from font(s) Arial.
plt.tight_layout()
C:\Users\user\AppData\Local\Temp\ipykernel_13356\850510082.py:55: UserWarning: Glyph 8322 (\N{SUBSCRIPT TWO}) missing from font(s) Arial.
plt.tight_layout()
C:\Users\user\AppData\Local\Temp\ipykernel_13356\850510082.py:55: UserWarning: Glyph 8323 (\N{SUBSCRIPT THREE}) missing from font(s) Arial.
plt.tight_layout()
C:\Users\user\AppData\Local\Temp\ipykernel_13356\850510082.py:55: UserWarning: Glyph 8477 (\N{DOUBLE-STRUCK CAPITAL R}) missing from font(s) Arial.
plt.tight_layout()
c:\Users\user\OneDrive\Documents\book\.venv\Lib\site-packages\IPython\core\pylabtools.py:170: UserWarning: Glyph 8321 (\N{SUBSCRIPT ONE}) missing from font(s) Arial.
fig.canvas.print_figure(bytes_io, **kw)
c:\Users\user\OneDrive\Documents\book\.venv\Lib\site-packages\IPython\core\pylabtools.py:170: UserWarning: Glyph 8322 (\N{SUBSCRIPT TWO}) missing from font(s) Arial.
fig.canvas.print_figure(bytes_io, **kw)
c:\Users\user\OneDrive\Documents\book\.venv\Lib\site-packages\IPython\core\pylabtools.py:170: UserWarning: Glyph 8323 (\N{SUBSCRIPT THREE}) missing from font(s) Arial.
fig.canvas.print_figure(bytes_io, **kw)
c:\Users\user\OneDrive\Documents\book\.venv\Lib\site-packages\IPython\core\pylabtools.py:170: UserWarning: Glyph 8477 (\N{DOUBLE-STRUCK CAPITAL R}) missing from font(s) Arial.
fig.canvas.print_figure(bytes_io, **kw)
4. Mathematical Formulation¶
Definition:
For S = {v₁, ..., vₖ} ⊆ V:
span(S) = { Σᵢ cᵢ vᵢ : cᵢ ∈ ℝ }Theorem: span(S) is a subspace of V.
Proof: (i) Take all cᵢ = 0 → 0 ∈ span(S). (ii) Sum of two linear combinations is a linear combination. (iii) Scalar multiple of a linear combination is a linear combination. ∎
Theorem: span(S) is the smallest subspace containing S.
Meaning: Any subspace W containing S must contain span(S).
In terms of matrices:
Let A = [v₁ | v₂ | ... | vₖ] (columns are the vectors)
span({v₁,...,vₖ}) = C(A) = {Ax : x ∈ ℝᵏ}
b ∈ span({v₁,...,vₖ}) ⟺ ∃x s.t. Ax = b ⟺ b ∈ C(A)Span does not increase when a dependent vector is added:
If vₖ₊₁ ∈ span({v₁,...,vₖ}), then span({v₁,...,vₖ,vₖ₊₁}) = span({v₁,...,vₖ})Dimension of the span:
dim(span({v₁,...,vₖ})) = rank([v₁ | ... | vₖ])The rank tells you how many of the vectors are actually contributing new directions. (Rank is formalized in ch157.)
5. Python Implementation¶
# --- Implementation: Span membership test and dimension ---
import numpy as np
def in_span(b, vectors, tol=1e-9):
"""
Test whether b is in the span of the given vectors.
Solves: A @ c = b (least squares) and checks the residual.
Args:
b: np.ndarray, shape (n,)
vectors: list of np.ndarray, each shape (n,)
tol: float — residual tolerance
Returns:
dict: {'in_span': bool, 'coefficients': np.ndarray or None, 'residual': float}
"""
A = np.column_stack(vectors)
c, _, _, _ = np.linalg.lstsq(A, b, rcond=None)
residual = float(np.linalg.norm(A @ c - b))
return {
'in_span': residual < tol,
'coefficients': c if residual < tol else None,
'residual': residual
}
def span_dimension(vectors, tol=1e-10):
"""
Compute the dimension of span(vectors).
Args:
vectors: list of np.ndarray
Returns:
int — rank of the matrix formed by columns
"""
A = np.column_stack(vectors)
return np.linalg.matrix_rank(A, tol=tol)
def minimal_spanning_set(vectors, tol=1e-10):
"""
Reduce a set of vectors to a minimal spanning set (a basis for their span).
Uses column pivoting via QR decomposition to identify independent columns.
Args:
vectors: list of np.ndarray
Returns:
list of np.ndarray — a subset of the input that spans the same space
"""
A = np.column_stack(vectors)
# QR with column pivoting: first k columns (after pivoting) are independent
Q, R, P = np.linalg.qr(A, mode='complete'), None, None
# Use SVD to identify independent columns more robustly
U, s, Vt = np.linalg.svd(A, full_matrices=False)
rank = np.sum(s > tol)
# Return U columns as the minimal spanning basis (orthonormalized)
return [U[:, i] for i in range(rank)]
# --- Tests ---
print("=== Span membership tests ===")
v1 = np.array([1.0, 0.0, 0.0])
v2 = np.array([0.0, 1.0, 0.0])
# b in the xy-plane: should be in span
b1 = np.array([3.0, -2.0, 0.0])
# b off the xy-plane: should NOT be in span
b2 = np.array([1.0, 1.0, 1.0])
r1 = in_span(b1, [v1, v2])
r2 = in_span(b2, [v1, v2])
print(f"b1={b1} in span({{e1,e2}}): {r1['in_span']} (coeffs: {r1['coefficients']})")
print(f"b2={b2} in span({{e1,e2}}): {r2['in_span']} (residual: {r2['residual']:.4f})")
print()
print("=== Span dimension ===")
# Three vectors in R^3 — but two are parallel
redundant = [
np.array([1.0, 2.0, 3.0]),
np.array([2.0, 4.0, 6.0]), # = 2 * first
np.array([0.0, 1.0, 0.0])
]
print(f"3 vectors (one redundant): span dim = {span_dimension(redundant)} (expected 2)")
independent = [
np.array([1.0, 0.0, 0.0]),
np.array([0.0, 1.0, 0.0]),
np.array([0.0, 0.0, 1.0])
]
print(f"3 independent vectors: span dim = {span_dimension(independent)} (expected 3)")=== Span membership tests ===
b1=[ 3. -2. 0.] in span({e1,e2}): True (coeffs: [ 3. -2.])
b2=[1. 1. 1.] in span({e1,e2}): False (residual: 1.0000)
=== Span dimension ===
3 vectors (one redundant): span dim = 2 (expected 2)
3 independent vectors: span dim = 3 (expected 3)
# --- Visualization: how span dimension changes as you add vectors ---
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-v0_8-whitegrid')
rng = np.random.default_rng(7)
AMBIENT_DIM = 5 # <-- try 3, 8, 10
MAX_VECTORS = 10 # number of vectors to add one by one
vectors = []
span_dims = []
for i in range(MAX_VECTORS):
# Alternate between truly new vectors and dependent ones
if i < AMBIENT_DIM:
# Independent: random vector unlikely to be in current span
v = rng.standard_normal(AMBIENT_DIM)
else:
# Dependent: linear combination of existing vectors
coeffs = rng.standard_normal(len(vectors))
v = sum(c * vec for c, vec in zip(coeffs, vectors))
vectors.append(v)
if len(vectors) == 1:
dim = 1
else:
dim = span_dimension(vectors)
span_dims.append(dim)
fig, ax = plt.subplots(figsize=(8, 4))
ax.plot(range(1, MAX_VECTORS+1), span_dims, 'bo-', markersize=8, linewidth=2)
ax.axhline(AMBIENT_DIM, color='red', linestyle='--', label=f'Max dim = {AMBIENT_DIM}')
ax.fill_between(range(1, MAX_VECTORS+1), span_dims,
alpha=0.15, color='blue')
ax.set_xlabel('Number of vectors added')
ax.set_ylabel('dim(span)')
ax.set_title(f'Span dimension grows until saturation at dim={AMBIENT_DIM}\n'
f'Adding dependent vectors (after index {AMBIENT_DIM}) changes nothing')
ax.set_xticks(range(1, MAX_VECTORS+1))
ax.legend()
plt.tight_layout()
plt.show()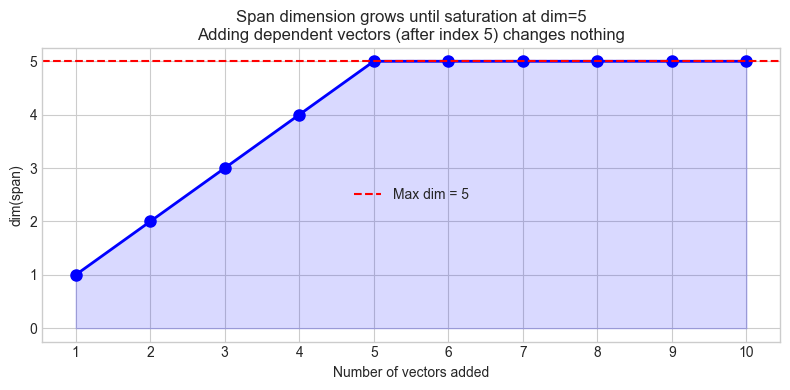
6. Experiments¶
# --- Experiment 1: When does a set of vectors span R^n? ---
# Hypothesis: k vectors can span R^n only if k >= n AND they are independent.
# Try changing: n and k to observe the threshold.
import numpy as np
rng = np.random.default_rng(42)
n = 4 # dimension of ambient space <-- modify
k_values = range(1, 8) # number of vectors to try <-- modify
print(f"Testing random vectors in R^{n}:")
print(f"{'k vectors':>12} {'dim(span)':>10} {'spans R^n?':>12}")
for k in k_values:
A = rng.standard_normal((n, k))
d = np.linalg.matrix_rank(A)
spans_all = (d == n)
print(f" k={k:2d} {d:10d} {str(spans_all):>12}")
print(f"\nConclusion: need at least k={n} linearly independent vectors to span R^{n}.")
print(f"With k > {n}, extra vectors are redundant (rank still capped at {n}).")Testing random vectors in R^4:
k vectors dim(span) spans R^n?
k= 1 1 False
k= 2 2 False
k= 3 3 False
k= 4 4 True
k= 5 4 True
k= 6 4 True
k= 7 4 True
Conclusion: need at least k=4 linearly independent vectors to span R^4.
With k > 4, extra vectors are redundant (rank still capped at 4).
# --- Experiment 2: Span as the column space of a matrix ---
# Hypothesis: b in span({v1,...,vk}) iff Ax=b has a solution.
# Try changing: the target b and observe when solutions exist.
import numpy as np
# Matrix A: columns are the spanning vectors
A = np.array([
[1, 0, 1],
[0, 1, 1],
[0, 0, 0] # third row zero: rank < 3
], dtype=float)
print(f"Matrix A (columns = spanning vectors):")
print(A)
print(f"Rank = {np.linalg.matrix_rank(A)} (spans a 2D subspace of R^3)")
print()
# b vectors to test
targets = [
(np.array([2.0, 3.0, 0.0]), "In column space (z=0)"),
(np.array([1.0, 1.0, 1.0]), "NOT in column space (z=1)"),
(np.array([0.5, -0.5, 0.0]), "In column space (z=0)"), # <-- modify
]
for b, description in targets:
c, residuals, rank, _ = np.linalg.lstsq(A, b, rcond=None)
residual = np.linalg.norm(A @ c - b)
in_cs = residual < 1e-9
print(f"b = {b} ({description})")
print(f" Residual: {residual:.2e} -> in span: {in_cs}")
if in_cs:
print(f" Coefficients: {c.round(4)}")
print()Matrix A (columns = spanning vectors):
[[1. 0. 1.]
[0. 1. 1.]
[0. 0. 0.]]
Rank = 2 (spans a 2D subspace of R^3)
b = [2. 3. 0.] (In column space (z=0))
Residual: 4.44e-16 -> in span: True
Coefficients: [0.3333 1.3333 1.6667]
b = [1. 1. 1.] (NOT in column space (z=1))
Residual: 1.00e+00 -> in span: False
b = [ 0.5 -0.5 0. ] (In column space (z=0))
Residual: 3.38e-16 -> in span: True
Coefficients: [ 0.5 -0.5 -0. ]
# --- Experiment 3: Removing vectors from a spanning set ---
# Hypothesis: A spanning set can be reduced to a basis without losing span.
# Approach: greedily remove vectors while maintaining the same span dimension.
import numpy as np
rng = np.random.default_rng(5)
N_DIM = 3 # ambient space <-- modify
N_VECTORS = 7 # start with an overcomplete set <-- modify
# Generate an overcomplete spanning set
base_vectors = [rng.standard_normal(N_DIM) for _ in range(N_DIM)]
extra_vectors = [sum(rng.random(N_DIM) * v for v in base_vectors)
for _ in range(N_VECTORS - N_DIM)]
all_vectors = base_vectors + extra_vectors
target_dim = np.linalg.matrix_rank(np.column_stack(all_vectors))
print(f"Starting with {len(all_vectors)} vectors, span dim = {target_dim}")
# Greedy reduction: try removing each vector; keep it removed if span dim unchanged
current_set = list(all_vectors)
removed = []
i = 0
while i < len(current_set):
trial = current_set[:i] + current_set[i+1:]
if len(trial) == 0:
break
d = np.linalg.matrix_rank(np.column_stack(trial))
if d == target_dim:
removed.append(i)
current_set = trial # remove it
else:
i += 1
final_dim = np.linalg.matrix_rank(np.column_stack(current_set))
print(f"After removal: {len(current_set)} vectors remain, span dim = {final_dim}")
print(f"This is a basis for the span: {len(current_set) == target_dim}")Starting with 7 vectors, span dim = 3
After removal: 3 vectors remain, span dim = 3
This is a basis for the span: True
7. Exercises¶
Easy 1. For each set, state the dimension of the span and describe it geometrically:
(a) {(1, 0, 0), (2, 0, 0)}
(b) {(1, 0, 0), (0, 1, 0)}
(c) {(1, 1, 0), (-1, 1, 0), (0, 0, 3)}
(Expected: 1 (a line), 2 (the xy-plane), 3 (all of ℝ³))
Easy 2. Write a function does_span_rn(vectors, n) that returns True if the vectors span ℝⁿ. Test it with several cases.
Medium 1. Given the vectors v₁ = (1, 2, 3), v₂ = (0, 1, 2), is b = (3, 7, 11) in span({v₁, v₂})? Find the coefficients or explain why not.
Medium 2. Given a 4×4 matrix A with rank 3, describe the span of its columns: what is its dimension, and in what space does it live? Now describe the span of the rows of A. Are they the same subspace? Same dimension?
Hard. Prove (computationally and in words): if span(S₁) = span(S₂) and both S₁ and S₂ are linearly independent, then |S₁| = |S₂| = dim(span). Write code to verify this for 50 random pairs of bases for the same subspace.
8. Mini Project¶
# --- Mini Project: Feature Reachability Analyzer ---
#
# Problem: In ML, a model can only represent targets that lie in the span
# of its features (for linear models). Given a dataset of features and targets,
# determine which targets are representable (in the column space of the
# feature matrix) and which are not.
#
# This is the fundamental question of linear regression: is y reachable?
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-v0_8-whitegrid')
rng = np.random.default_rng(0)
# --- Dataset ---
# Features: x1 (income), x2 (age) — 2 features, N samples
N = 50
x1 = rng.uniform(20, 100, N) # income (thousands)
x2 = rng.uniform(20, 65, N) # age
X = np.column_stack([x1, x2]) # feature matrix: shape (N, 2)
# Three target vectors:
# y1: perfectly representable (linear combo of features)
y1 = 0.5*x1 - 0.3*x2 + 0.0 # exact linear combo — in span
# y2: representable with noise
y2 = 0.5*x1 - 0.3*x2 + rng.normal(0, 2, N) # noisy — approximately in span
# y3: not representable (depends on x1^2)
y3 = 0.01 * x1**2 + rng.normal(0, 1, N) # nonlinear — NOT in span of linear features
def fit_and_residual(X, y):
"""
Fit y to span of X (least squares) and return residual norm.
Large residual = y not in span(X).
"""
coeffs, _, _, _ = np.linalg.lstsq(X, y, rcond=None)
y_hat = X @ coeffs
residual = np.linalg.norm(y - y_hat)
r2 = 1 - np.var(y - y_hat) / np.var(y)
return coeffs, y_hat, residual, r2
# --- TODO: run analysis for each target ---
targets = [('y1 (exact linear)', y1), ('y2 (linear + noise)', y2), ('y3 (nonlinear)', y3)]
fig, axes = plt.subplots(1, 3, figsize=(13, 4))
for ax, (name, y) in zip(axes, targets):
coeffs, y_hat, residual, r2 = fit_and_residual(X, y)
ax.scatter(y, y_hat, alpha=0.5, s=20)
mn, mx = min(y.min(), y_hat.min()), max(y.max(), y_hat.max())
ax.plot([mn, mx], [mn, mx], 'r--', lw=1.5, label='Perfect fit')
ax.set_xlabel('Actual y')
ax.set_ylabel('Predicted ŷ')
ax.set_title(f'{name}\nR²={r2:.3f} residual={residual:.2f}')
ax.legend(fontsize=8)
print(f"{name}: R²={r2:.4f}, residual norm={residual:.4f}")
print(f" Coefficients: {coeffs.round(4)}")
in_span = r2 > 0.99
print(f" In span of X: {in_span}")
print()
plt.suptitle('Feature Reachability: Is the target in span(X)?', fontsize=12)
plt.tight_layout()
plt.show()9. Chapter Summary & Connections¶
What was covered:
The span of a set S is the set of all linear combinations — every vector reachable from S.
Span is always a subspace; it is the smallest subspace containing S.
Adding a dependent vector to a spanning set does not change the span.
Span dimension = rank of the matrix formed by the vectors.
Testing span membership reduces to solving a linear system.
Backward connection: Span connects ch127 (Linear Combination) (introduced in ch127) with ch138–139 (Subspaces, Basis): the span of the basis vectors is the entire space, and that is precisely the definition of a basis — a minimal spanning set.
Forward connections:
In ch141 (Linear Independence), we examine what makes a set of vectors not redundant — the condition that ensures each vector expands the span.
This reappears in ch160 (Systems of Linear Equations): the question “does Ax = b have a solution?” is exactly “is b in the column space (span of columns) of A?”
In ch186 (PCA), the span of the top-k principal components is the k-dimensional subspace that captures the most variance in data — span becomes the language of dimensionality reduction.