Prerequisites: ch157 (Matrix Inverse), ch160 (Systems of Linear Equations), ch173 (SVD), ch192 (Rank, Nullity) You will learn:
What the Moore-Penrose pseudoinverse computes
Why it solves over- and under-determined systems optimally
How to compute it via SVD
The four Moore-Penrose conditions and why they matter
Connection to linear regression Environment: Python 3.x, numpy, matplotlib
1. Concept¶
The inverse only exists when is square and full rank. In applications, is almost never square — we have more equations than unknowns (overdetermined) or fewer (underdetermined). The Moore-Penrose pseudoinverse generalizes the inverse to handle both cases:
Overdetermined (): gives the least-squares solution — minimizes
Underdetermined (): gives the minimum-norm solution — minimizes among all solutions
Square, full rank: exactly
Common misconception: does not solve exactly in the overdetermined case. It finds the that makes closest to in L2 norm.
2. Intuition & Mental Models¶
Geometric (overdetermined): You have noisy measurements of an -dimensional quantity. cannot exactly equal every measurement simultaneously. finds the whose prediction is closest to the data — the orthogonal projection of onto the column space of .
Geometric (underdetermined): Infinitely many satisfy . They form an affine subspace. is the point in that subspace closest to the origin — it uses the minimum amount of “resource” to explain the data.
Computational: Via SVD , invert only the non-zero singular values: where replaces each non-zero with . This is safe because it never divides by zero (ch173 — SVD).
3. Visualization¶
# --- Visualization: Pseudoinverse for overdetermined system (linear fit) ---
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-v0_8-whitegrid')
rng = np.random.default_rng(42)
# Overdetermined: fit y = a + b*x to noisy data (10 points, 2 parameters)
N_PTS = 15
x_data = np.linspace(0, 3, N_PTS)
y_true = 2.0 + 1.5 * x_data
y_noisy = y_true + rng.normal(0, 0.5, N_PTS)
# Design matrix: each row is [1, x_i]
A = np.column_stack([np.ones(N_PTS), x_data]) # (15, 2)
# Pseudoinverse solution
def pseudoinverse(A, tol=1e-12):
"""Moore-Penrose pseudoinverse via SVD."""
U, s, Vt = np.linalg.svd(A, full_matrices=False)
s_inv = np.where(np.abs(s) > tol, 1.0/s, 0.0)
return Vt.T @ np.diag(s_inv) @ U.T
A_pinv = pseudoinverse(A)
x_ls = A_pinv @ y_noisy # least-squares solution
# Residuals
y_fit = A @ x_ls
residuals = y_noisy - y_fit
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
# Scatter + fit line
axes[0].scatter(x_data, y_noisy, color='steelblue', s=50, label='Observations', zorder=3)
x_line = np.linspace(0, 3, 100)
axes[0].plot(x_line, x_ls[0] + x_ls[1]*x_line, 'r-', lw=2, label=f'Fit: y={x_ls[0]:.2f}+{x_ls[1]:.2f}x')
axes[0].plot(x_line, 2 + 1.5*x_line, 'g--', lw=1.5, alpha=0.7, label='True: y=2+1.5x')
for xi, yi, yf in zip(x_data, y_noisy, y_fit):
axes[0].plot([xi, xi], [yi, yf], 'gray', lw=0.8, alpha=0.5)
axes[0].set_xlabel('x')
axes[0].set_ylabel('y')
axes[0].set_title('Least-Squares Fit via Pseudoinverse')
axes[0].legend(fontsize=8)
# Residual plot
axes[1].bar(range(N_PTS), residuals, color='steelblue', alpha=0.8)
axes[1].axhline(0, color='black', lw=1)
axes[1].set_xlabel('Data point index')
axes[1].set_ylabel('Residual (y - y_fit)')
axes[1].set_title(f'Residuals (||Ax-b||² = {np.sum(residuals**2):.4f})')
plt.tight_layout()
plt.show()
print(f"Least-squares solution: a={x_ls[0]:.4f}, b={x_ls[1]:.4f}")
print(f"True parameters: a=2.0000, b=1.5000")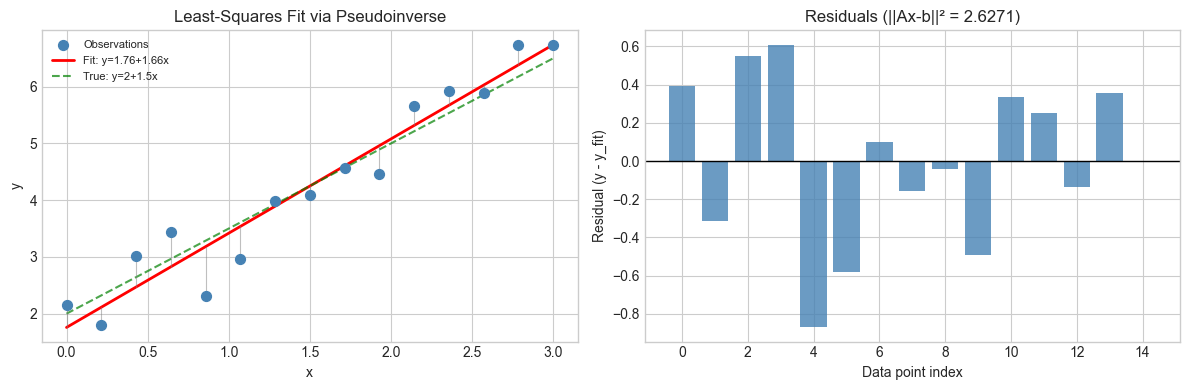
Least-squares solution: a=1.7573, b=1.6610
True parameters: a=2.0000, b=1.5000
4. Mathematical Formulation¶
Given with SVD :
where replaces each non-zero with (and keeps zeros as zero).
The four Moore-Penrose conditions characterize uniquely:
(weak inverse)
(reflexive)
(Hermitian projection onto col)
(Hermitian projection onto row)
Normal equations (overdetermined case): the least-squares solution satisfies . When is invertible, .
5. Python Implementation¶
# --- Implementation: Pseudoinverse and verification of Moore-Penrose conditions ---
def pseudoinverse_svd(A, tol=1e-12):
"""
Moore-Penrose pseudoinverse via SVD.
A+ = V @ diag(1/s) @ U.T (only inverting non-zero singular values)
"""
U, s, Vt = np.linalg.svd(A, full_matrices=False)
s_inv = np.where(np.abs(s) > tol, 1.0 / s, 0.0)
return Vt.T @ np.diag(s_inv) @ U.T
def verify_moore_penrose(A, A_plus, tol=1e-8):
"""Verify the four Moore-Penrose conditions."""
cond1 = np.linalg.norm(A @ A_plus @ A - A) # A A+ A = A
cond2 = np.linalg.norm(A_plus @ A @ A_plus - A_plus) # A+ A A+ = A+
cond3 = np.linalg.norm((A @ A_plus) - (A @ A_plus).T) # (A A+)' = A A+
cond4 = np.linalg.norm((A_plus @ A) - (A_plus @ A).T) # (A+ A)' = A+ A
return {'AA+A=A': cond1, 'A+AA+=A+': cond2,
'(AA+)sym': cond3, '(A+A)sym': cond4}
# Test on several matrix shapes
test_cases = [
('Overdetermined 6x3 rank-3', rng.normal(0,1,(6,3))),
('Underdetermined 3x6 rank-3', rng.normal(0,1,(3,6))),
('Square full rank 4x4', rng.normal(0,1,(4,4))),
('Rank-deficient 4x5 rank-2', np.outer(rng.normal(0,1,4), rng.normal(0,1,5)) +
np.outer(rng.normal(0,1,4), rng.normal(0,1,5))),
]
for name, A_t in test_cases:
Ap = pseudoinverse_svd(A_t)
errs = verify_moore_penrose(A_t, Ap)
all_ok = all(v < 1e-8 for v in errs.values())
max_err = max(errs.values())
print(f"{name}: {'PASS' if all_ok else 'FAIL'} (max error={max_err:.2e})")Overdetermined 6x3 rank-3: PASS (max error=1.13e-15)
Underdetermined 3x6 rank-3: PASS (max error=1.38e-15)
Square full rank 4x4: PASS (max error=1.19e-15)
Rank-deficient 4x5 rank-2: PASS (max error=9.22e-16)
6. Experiments¶
# --- Experiment: Minimum-norm solution for underdetermined system ---
# Hypothesis: A+b gives the solution with smallest ||x|| among all exact solutions.
# Try changing: N_VARS, N_EQS
N_EQS = 3 # <-- fewer equations than variables
N_VARS = 7 # <-- try 5, 7, 10
A_under = rng.normal(0, 1, (N_EQS, N_VARS))
b_under = rng.normal(0, 1, N_EQS)
# Pseudoinverse (minimum norm) solution
x_minnorm = pseudoinverse_svd(A_under) @ b_under
# Sample other solutions: x_minnorm + alpha * null_vector
_, ns, _, _, _ = __import__('__main__').__dict__.get(
'four_fundamental_subspaces', lambda A: (None, np.zeros((N_VARS,0)), None, None, None))(A_under)
# Recompute null space
_, _, Vt_u = np.linalg.svd(A_under, full_matrices=True)
null_basis = Vt_u[N_EQS:, :].T # null space basis vectors
# Sample norms of random solutions
norms_sampled = []
for _ in range(200):
alpha = rng.normal(0, 1, null_basis.shape[1])
x_rand = x_minnorm + null_basis @ alpha
norms_sampled.append(np.linalg.norm(x_rand))
fig, ax = plt.subplots(figsize=(8, 3))
ax.hist(norms_sampled, bins=30, color='steelblue', alpha=0.8, label='Random solutions')
ax.axvline(np.linalg.norm(x_minnorm), color='crimson', lw=2.5,
label=f'Pseudoinverse: ||x||={np.linalg.norm(x_minnorm):.3f}')
ax.set_xlabel('||x|| (norm of solution)')
ax.set_ylabel('Count')
ax.set_title(f'Minimum-Norm Solution ({N_EQS} eqs, {N_VARS} vars)')
ax.legend(fontsize=8)
plt.tight_layout()
plt.show()
print(f"Pseudoinverse gives the minimum-norm solution: {np.linalg.norm(x_minnorm):.4f}")
print(f"All sampled solutions have norm >= {np.linalg.norm(x_minnorm):.4f}")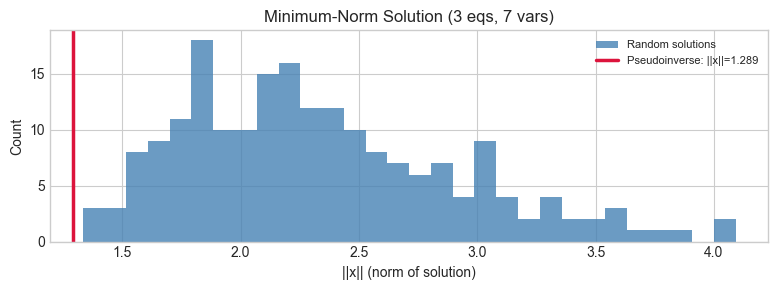
Pseudoinverse gives the minimum-norm solution: 1.2895
All sampled solutions have norm >= 1.2895
7. Exercises¶
Easy 1. Compute the pseudoinverse of a row vector . What does compute geometrically? (Hint: it projects onto the span of )
Easy 2. Show numerically that for an invertible square matrix, pseudoinverse_svd(A) ≈ np.linalg.inv(A) by checking their element-wise difference.
Medium 1. The normal equations can be solved directly. Implement both methods (normal equations via np.linalg.solve and pseudoinverse via SVD) and compare numerical stability by conditioning to have a large condition number (). Which approach is more stable?
Medium 2. Implement polynomial curve fitting using the pseudoinverse. Given data points , fit a degree- polynomial by building the Vandermonde design matrix and computing . Show results for on noisy sinusoidal data.
Hard. The Tikhonov regularized pseudoinverse replaces with . Derive analytically why this is preferable when has small singular values. Implement it via SVD () and show how controls the norm-accuracy tradeoff.
8. Mini Project¶
# --- Mini Project: Image Deblurring via Pseudoinverse ---
# Problem: A blurred image satisfies y = A x where A is a known blur operator.
# Recover x from y using the pseudoinverse.
# Dataset: Synthetic 1D signal with a known convolution blur.
# 1D signal
N_SIGNAL = 60
x_true = np.zeros(N_SIGNAL)
for pos in [10, 25, 40, 50]:
x_true[pos:pos+5] = 1.0 # rectangular pulses
# Blur matrix: convolution with a Gaussian kernel
BLUR_WIDTH = 3 # <-- try 2, 3, 5
kernel = np.exp(-np.arange(-BLUR_WIDTH, BLUR_WIDTH+1)**2 / (2*BLUR_WIDTH**2))
kernel /= kernel.sum()
# Build circulant convolution matrix A
A_blur = np.zeros((N_SIGNAL, N_SIGNAL))
for i in range(N_SIGNAL):
for ki, kv in enumerate(kernel):
j = (i + ki - BLUR_WIDTH) % N_SIGNAL
A_blur[i, j] += kv
y_blurred = A_blur @ x_true + rng.normal(0, 0.02, N_SIGNAL)
# Naive pseudoinverse (amplifies noise for ill-conditioned A)
x_recovered_naive = pseudoinverse_svd(A_blur, tol=1e-12) @ y_blurred
# Regularized (Tikhonov): replace s_i -> s_i / (s_i^2 + lam)
LAM = 0.01 # <-- try 0.001, 0.01, 0.1
U_b, s_b, Vt_b = np.linalg.svd(A_blur, full_matrices=False)
s_reg = s_b / (s_b**2 + LAM)
A_reg_inv = Vt_b.T @ np.diag(s_reg) @ U_b.T
x_recovered_reg = A_reg_inv @ y_blurred
fig, axes = plt.subplots(4, 1, figsize=(12, 9), sharex=True)
axes[0].plot(x_true, 'steelblue', lw=2, label='True signal')
axes[0].set_title('True Signal'); axes[0].set_ylabel('Amplitude')
axes[1].plot(y_blurred, 'darkorange', lw=1.5, label='Blurred + noise')
axes[1].set_title('Blurred + Noise'); axes[1].set_ylabel('Amplitude')
axes[2].plot(x_recovered_naive, 'crimson', lw=1, label='Naive pseudoinverse')
axes[2].set_title('Recovered (naive pseudoinverse — may amplify noise)')
axes[2].set_ylabel('Amplitude')
axes[3].plot(x_recovered_reg, 'forestgreen', lw=1.5, label=f'Tikhonov (λ={LAM})')
axes[3].set_title(f'Recovered (Tikhonov λ={LAM})')
axes[3].set_xlabel('Index')
axes[3].set_ylabel('Amplitude')
plt.tight_layout()
plt.show()
err_naive = np.linalg.norm(x_recovered_naive - x_true)
err_reg = np.linalg.norm(x_recovered_reg - x_true)
print(f"Recovery error — Naive: {err_naive:.4f}, Tikhonov: {err_reg:.4f}")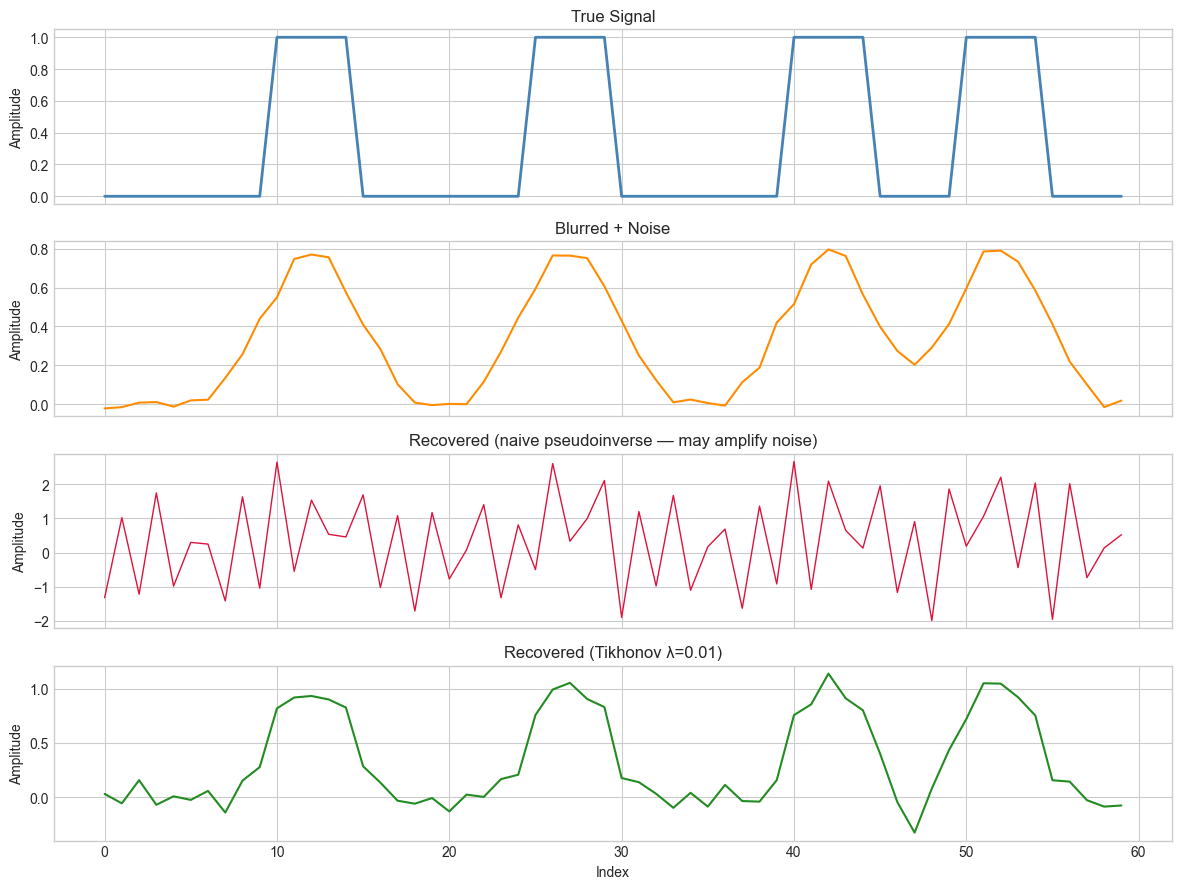
Recovery error — Naive: 9.6685, Tikhonov: 1.2110
9. Chapter Summary & Connections¶
The Moore-Penrose pseudoinverse generalizes matrix inversion to non-square and rank-deficient matrices (ch173 — SVD).
For overdetermined systems: minimizes (least squares). For underdetermined: minimizes among all exact solutions.
Small singular values make the pseudoinverse ill-conditioned; Tikhonov regularization () damps them and stabilizes the solution at the cost of some accuracy.
Forward: This reappears in ch182 (Linear Regression via Matrix Algebra) as the normal equations, in ch212 (Gradient Descent) as the closed-form solution of the quadratic loss, and in ch273 (Regression) as the sklearn LinearRegression estimator’s internal solver. Tikhonov regularization is Ridge regression (ch273).
Backward: This chapter synthesizes ch157 (inverse), ch160 (systems), ch173 (SVD), and ch192 (rank/nullity). The pseudoinverse is the answer to “what does matrix inversion mean when the matrix isn’t square?”