Prerequisites: ch128 (Vector Length/Norm), ch173 (SVD), ch160 (Systems of Linear Equations), ch193 (Pseudoinverse) You will learn:
The Frobenius norm, spectral norm, and nuclear norm
How to measure the “size” of a linear transformation
The condition number κ(A) and why it matters for numerical stability
Why ill-conditioned matrices produce unreliable solutions
How to detect and mitigate numerical ill-conditioning Environment: Python 3.x, numpy, matplotlib
1. Concept¶
A vector norm measures the size of a vector. A matrix norm measures the size of a linear transformation — how much it can stretch or shrink vectors. There are many valid norms; each captures a different aspect of matrix behavior.
The condition number measures how sensitive the solution of is to small perturbations in or . A large means small changes in data produce large changes in solution — numerically dangerous.
Common misconception: an ill-conditioned matrix is not the same as a singular matrix. Ill-conditioning is a continuous spectrum — a matrix can be nearly-singular (very large ) without being exactly singular.
2. Intuition & Mental Models¶
Geometric: A matrix transforms the unit sphere into an ellipsoid. The largest axis of the ellipsoid has length (largest singular value); the smallest has length . The spectral norm is ; the condition number is .
Numerical stability: If , you lose approximately digits of accuracy when solving in floating point. A condition number of 1012 on a 64-bit system (about 15 significant digits) leaves only 3 digits of accuracy.
Think of it as: the condition number is the ratio of the most to least responsive direction. A well-conditioned matrix responds similarly in all directions; an ill-conditioned one is nearly flat in some direction.
3. Visualization¶
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-v0_8-whitegrid')
rng = np.random.default_rng(42)
# Visualize how a matrix transforms the unit circle
def transform_unit_circle(A, n_pts=200):
theta = np.linspace(0, 2*np.pi, n_pts)
circle = np.array([np.cos(theta), np.sin(theta)]) # (2, n_pts)
ellipse = A @ circle # (2, n_pts)
return circle, ellipse
# Three 2x2 matrices with different condition numbers
kappa_vals = [1.0, 10.0, 100.0]
matrices = []
for kappa in kappa_vals:
# Build matrix with singular values [kappa, 1] rotated 30 degrees
angle = np.pi / 6
R = np.array([[np.cos(angle), -np.sin(angle)], [np.sin(angle), np.cos(angle)]])
S = np.diag([kappa, 1.0])
matrices.append(R @ S @ R.T)
fig, axes = plt.subplots(1, 3, figsize=(13, 4))
for ax, A, kappa in zip(axes, matrices, kappa_vals):
circle, ellipse = transform_unit_circle(A)
ax.plot(circle[0], circle[1], 'b--', lw=1, alpha=0.5, label='Unit circle')
ax.plot(ellipse[0], ellipse[1], 'r-', lw=2, label='Transformed')
sigma1, sigma2 = np.linalg.svd(A, compute_uv=False)[:2]
ax.set_title(f'κ={kappa:.0f} (σ₁={sigma1:.1f}, σ₂={sigma2:.1f})', fontsize=9)
ax.set_aspect('equal')
ax.legend(fontsize=7)
ax.axhline(0, color='black', lw=0.5)
ax.axvline(0, color='black', lw=0.5)
plt.suptitle('Unit Circle Transformed by Matrices with Different Condition Numbers', fontsize=10)
plt.tight_layout()
plt.show()
print('Higher κ → more eccentric ellipse → larger ratio of max to min stretching.')C:\Users\user\AppData\Local\Temp\ipykernel_10588\722168776.py:36: UserWarning: Glyph 8321 (\N{SUBSCRIPT ONE}) missing from font(s) Arial.
plt.tight_layout()
C:\Users\user\AppData\Local\Temp\ipykernel_10588\722168776.py:36: UserWarning: Glyph 8322 (\N{SUBSCRIPT TWO}) missing from font(s) Arial.
plt.tight_layout()
c:\Users\user\OneDrive\Documents\book\.venv\Lib\site-packages\IPython\core\pylabtools.py:170: UserWarning: Glyph 8321 (\N{SUBSCRIPT ONE}) missing from font(s) Arial.
fig.canvas.print_figure(bytes_io, **kw)
c:\Users\user\OneDrive\Documents\book\.venv\Lib\site-packages\IPython\core\pylabtools.py:170: UserWarning: Glyph 8322 (\N{SUBSCRIPT TWO}) missing from font(s) Arial.
fig.canvas.print_figure(bytes_io, **kw)
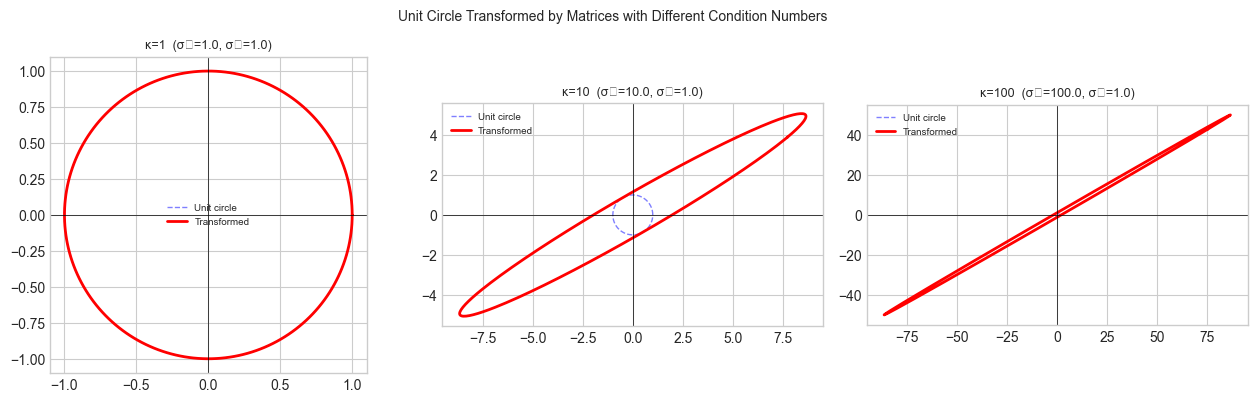
Higher κ → more eccentric ellipse → larger ratio of max to min stretching.
4. Mathematical Formulation¶
Common matrix norms for with singular values :
Condition number (spectral):
Perturbation bound: if , then:
The condition number is the worst-case amplification factor for relative errors.
5. Python Implementation¶
def matrix_norms(A):
"""
Compute Frobenius, spectral (L2), and nuclear norms.
Args: A: (m, n) matrix
Returns: dict of norm values
"""
_, s, _ = np.linalg.svd(A, full_matrices=False)
return {
'frobenius': np.sqrt(np.sum(A**2)), # sqrt(sum of squares)
'frobenius_svd': np.sqrt(np.sum(s**2)), # same via SVD
'spectral': s[0], # largest singular value
'nuclear': s.sum(), # sum of singular values
}
def condition_number(A, tol=1e-15):
"""
Spectral condition number κ₂(A) = σ_max / σ_min.
Returns np.inf for singular/near-singular matrices.
"""
_, s, _ = np.linalg.svd(A, full_matrices=False)
if s[-1] < tol:
return np.inf
return s[0] / s[-1]
# Build a family of matrices with controlled condition numbers
for target_kappa in [1, 10, 100, 1000, 1e6]:
# Use SVD to build a matrix with exactly this condition number
U, _, Vt = np.linalg.svd(rng.normal(0, 1, (4, 4)), full_matrices=False)
s_controlled = np.array([target_kappa, np.sqrt(target_kappa), np.sqrt(target_kappa), 1.0])
A_test = U @ np.diag(s_controlled) @ Vt
kappa_computed = condition_number(A_test)
norms = matrix_norms(A_test)
print(f'κ_target={target_kappa:.0e} κ_computed={kappa_computed:.4e} ||A||_F={norms["frobenius"]:.3f} ||A||_2={norms["spectral"]:.3f}')κ_target=1e+00 κ_computed=1.0000e+00 ||A||_F=2.000 ||A||_2=1.000
κ_target=1e+01 κ_computed=1.0000e+01 ||A||_F=11.000 ||A||_2=10.000
κ_target=1e+02 κ_computed=1.0000e+02 ||A||_F=101.000 ||A||_2=100.000
κ_target=1e+03 κ_computed=1.0000e+03 ||A||_F=1001.000 ||A||_2=1000.000
κ_target=1e+06 κ_computed=1.0000e+06 ||A||_F=1000001.000 ||A||_2=1000000.000
6. Experiments¶
# --- Experiment: Condition number vs numerical solution error ---
# Hypothesis: higher κ leads to larger solution error for perturbed RHS.
# Try changing: KAPPA_RANGE, NOISE_LEVEL
KAPPA_RANGE = [1, 5, 20, 100, 500, 2000, 1e5]
NOISE_LEVEL = 1e-4 # <-- try 1e-3, 1e-5
errors = []
for kappa in KAPPA_RANGE:
# Build 4x4 matrix with condition number kappa
U_e, _, Vt_e = np.linalg.svd(rng.normal(0, 1, (4, 4)), full_matrices=False)
s_e = np.array([kappa, kappa**(2/3), kappa**(1/3), 1.0])
A_e = U_e @ np.diag(s_e) @ Vt_e
x_exact = rng.normal(0, 1, 4)
b_exact = A_e @ x_exact
b_noisy = b_exact + NOISE_LEVEL * rng.normal(0, 1, 4)
x_exact_sol = np.linalg.solve(A_e, b_exact)
x_noisy_sol = np.linalg.solve(A_e, b_noisy)
rel_err = np.linalg.norm(x_noisy_sol - x_exact_sol) / np.linalg.norm(x_exact_sol)
errors.append(rel_err)
fig, ax = plt.subplots(figsize=(8, 4))
ax.loglog(KAPPA_RANGE, errors, 'o-', ms=6, color='steelblue', label='Actual error')
ax.loglog(KAPPA_RANGE, [NOISE_LEVEL * k for k in KAPPA_RANGE], 'r--', lw=1.5, label='κ × noise (bound)')
ax.set_xlabel('Condition number κ')
ax.set_ylabel('Relative solution error')
ax.set_title(f'Condition Number vs Solution Error (noise={NOISE_LEVEL:.0e})')
ax.legend()
plt.tight_layout()
plt.show()
print('Error grows proportionally with κ — the condition number is a sensitivity multiplier.')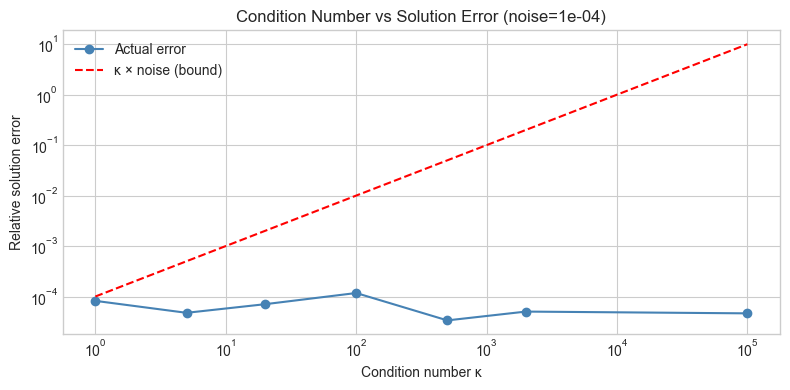
Error grows proportionally with κ — the condition number is a sensitivity multiplier.
7. Exercises¶
Easy 1. For a identity matrix, compute all three norms (Frobenius, spectral, nuclear). What are they? Why?
Easy 2. The Hilbert matrix is famously ill-conditioned. Compute its condition number for sizes and plot vs .
Medium 1. Implement a function that builds an matrix with a specified condition number using SVD (place singular values at geometric spacing from 1 to ). Verify by computing the condition number of the result.
Medium 2. Show that where . Verify this inequality for 100 random matrices of varying shape and rank.
Hard. The nuclear norm is convex in . It is the convex relaxation of rank (since rank counts non-zeros and sums them). Implement nuclear norm minimization via iterative soft-thresholding of singular values: . Apply this to low-rank matrix completion on a partially-observed matrix.
8. Mini Project¶
# --- Mini Project: Numerical Stability Benchmark ---
# Compare three approaches to solving Ax=b on matrices with increasing κ.
def solve_normal_equations(A, b):
"""(A.T A)^{-1} A.T b — numerically unstable for large κ."""
return np.linalg.solve(A.T @ A, A.T @ b)
def solve_qr(A, b):
"""QR decomposition: A = QR, x = R^{-1} Q.T b — more stable."""
Q, R = np.linalg.qr(A)
return np.linalg.solve(R, Q.T @ b)
def solve_svd(A, b, tol=1e-12):
"""Pseudoinverse via SVD — most stable."""
U, s, Vt = np.linalg.svd(A, full_matrices=False)
s_inv = np.where(s > tol, 1.0/s, 0.0)
return Vt.T @ (s_inv * (U.T @ b))
kappas = [1, 10, 100, 1000, 1e5, 1e8, 1e10]
m, n = 20, 5
errs_ne, errs_qr, errs_svd = [], [], []
for kappa in kappas:
U_b, _, Vt_b = np.linalg.svd(rng.normal(0, 1, (m, n)), full_matrices=False)
s_b = np.geomspace(kappa, 1, n)
A_b = U_b @ np.diag(s_b) @ Vt_b
x_true = rng.normal(0, 1, n)
b_b = A_b @ x_true
for solver, errs in [(solve_normal_equations, errs_ne),
(solve_qr, errs_qr),
(solve_svd, errs_svd)]:
try:
x_sol = solver(A_b, b_b)
errs.append(np.linalg.norm(x_sol - x_true) / np.linalg.norm(x_true))
except Exception:
errs.append(np.nan)
fig, ax = plt.subplots(figsize=(9, 4))
ax.loglog(kappas, errs_ne, 'o-', ms=5, color='crimson', label='Normal equations')
ax.loglog(kappas, errs_qr, 's-', ms=5, color='steelblue', label='QR decomposition')
ax.loglog(kappas, errs_svd, '^-', ms=5, color='forestgreen', label='SVD pseudoinverse')
ax.axhline(1e-15, color='black', ls=':', lw=1, label='Machine precision')
ax.set_xlabel('Condition number κ')
ax.set_ylabel('Relative solution error')
ax.set_title('Numerical Stability: Normal Equations vs QR vs SVD')
ax.legend(fontsize=8)
plt.tight_layout()
plt.show()
print('Normal equations square the condition number (κ(A.T A) = κ(A)²).')
print('QR and SVD work at κ rather than κ², giving more reliable solutions.')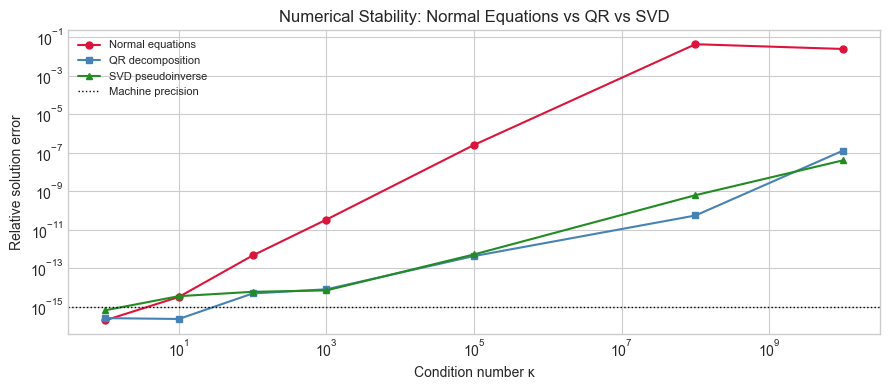
Normal equations square the condition number (κ(A.T A) = κ(A)²).
QR and SVD work at κ rather than κ², giving more reliable solutions.
9. Chapter Summary & Connections¶
Matrix norms measure the size of linear maps: Frobenius = RMS of all entries; spectral = max singular value; nuclear = sum of singular values (ch173).
The condition number measures numerical sensitivity: a relative perturbation of size in can produce a relative error of up to in .
Squaring the matrix via normal equations squares the condition number. Prefer QR or SVD for numerical linear algebra.
Nuclear norm minimization is the standard convex relaxation of rank-minimization, used in matrix completion and low-rank recovery problems.
Forward: Condition numbers reappear in ch212 (Gradient Descent) as the convergence rate of gradient descent on a quadratic depends on where is the Hessian. Ill-conditioned quadratics require many more iterations. Nuclear norm appears in ch289 (collaborative filtering) as a regularizer for low-rank matrix completion.
Backward: This chapter makes precise the numerical warnings in ch179 (Numerical Linear Algebra in Practice), which cautioned against taking explicit inverses. The relationship depends entirely on ch173 (SVD).