Prerequisites: ch151 (Introduction to Matrices), ch154 (Matrix Multiplication), ch160 (Systems of Linear Equations), ch192 (Rank, Nullity) You will learn:
What makes a matrix sparse and why it matters computationally
COO, CSR, and CSC storage formats
Sparse matrix-vector multiplication from scratch
How sparsity arises in real applications (graphs, finite elements, NLP)
The tradeoff between storage format and operation efficiency Environment: Python 3.x, numpy, matplotlib
1. Concept¶
A sparse matrix is one where most entries are zero. The standard dense representation wastes both memory and computation on these zeros. For a matrix with 10 non-zeros per row, dense storage requires 1012 floats (8 TB); sparse storage requires only 107 floats (80 MB).
Sparsity arises wherever structure limits interaction:
Graphs: adjacency matrix of a sparse graph
Finite element methods: each element touches only nearby elements
NLP: term-document matrix (most words don’t appear in most documents)
Neural networks: convolutional layers have sparse connectivity patterns
Common misconception: sparse formats are not always faster. For dense operations (e.g., matrix-matrix product of two sparse matrices producing a dense result), dense storage can win.
2. Intuition & Mental Models¶
Think of a sparse matrix as a list of non-zero entries, not a grid. Instead of storing 1 million zeros, store 10,000 (row, col, value) triples.
Three formats, each optimized for different operations:
COO (Coordinate):
(rows, cols, vals)— easy to construct, slow for arithmeticCSR (Compressed Sparse Row): row pointers + column indices — fast row access and matrix-vector product
CSC (Compressed Sparse Column): column pointers + row indices — fast column access and transpose operations
Recall from ch154 (Matrix Multiplication): dense takes operations. Sparse takes where = number of non-zeros. If , this is a massive speedup.
3. Visualization¶
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-v0_8-whitegrid')
rng = np.random.default_rng(42)
# Visualize sparsity patterns of matrices from different domains
def random_sparse(n, nnz_per_row, rng=rng):
"""Random sparse matrix with fixed nnz per row."""
rows, cols, vals = [], [], []
for i in range(n):
js = rng.choice(n, nnz_per_row, replace=False)
for j in js:
rows.append(i)
cols.append(j)
vals.append(rng.normal())
return np.array(rows), np.array(cols), np.array(vals), (n, n)
def tridiagonal_matrix(n):
"""Tridiagonal matrix (finite difference Laplacian)."""
rows, cols, vals = [], [], []
for i in range(n):
rows.append(i); cols.append(i); vals.append(2.0)
if i > 0: rows.append(i); cols.append(i-1); vals.append(-1.0)
if i < n-1: rows.append(i); cols.append(i+1); vals.append(-1.0)
return np.array(rows), np.array(cols), np.array(vals), (n, n)
def sparse_to_dense(rows, cols, vals, shape):
A = np.zeros(shape)
for r, c, v in zip(rows, cols, vals):
A[r, c] += v
return A
n_vis = 40
fig, axes = plt.subplots(1, 3, figsize=(13, 4))
for ax, (rows, cols, vals, shape), title in [
(axes[0], random_sparse(n_vis, 3), 'Random sparse (3 nnz/row)'),
(axes[1], tridiagonal_matrix(n_vis), 'Tridiagonal (1D Laplacian)'),
(axes[2], random_sparse(n_vis, 10), 'Random sparse (10 nnz/row)'),
]:
A_dense = sparse_to_dense(rows, cols, vals, shape)
ax.spy(A_dense, markersize=3)
nnz = len(vals)
sparsity = 1 - nnz / (n_vis**2)
ax.set_title(f'{title}\nnnz={nnz}, sparsity={sparsity:.1%}', fontsize=8)
plt.suptitle('Sparsity Patterns (spy plots)', fontsize=10)
plt.tight_layout()
plt.show()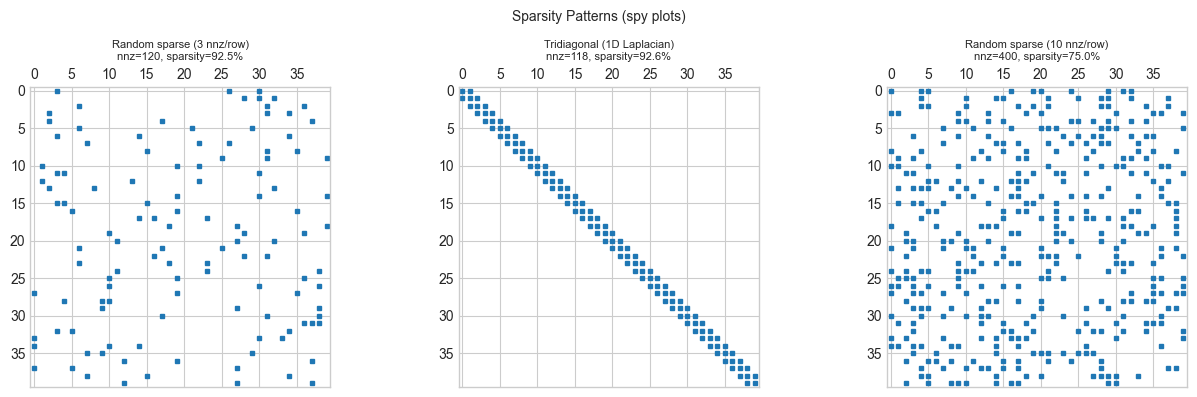
4. Mathematical Formulation¶
COO format for with non-zeros:
Storage: numbers. Access to : scan.
CSR format adds a row pointer array indptr of length :
indptr[i]= index indata/col_idxwhere row startsindptr[i+1] - indptr[i]= number of non-zeros in row
Storage: numbers. Row access: . Matrix-vector product:
Time: total — one pass through all non-zeros.
5. Python Implementation¶
# --- Sparse Matrix Classes: COO and CSR ---
class SparseMatrixCOO:
"""
COO (Coordinate) sparse matrix.
Stores triplets (row, col, val) for each non-zero.
"""
def __init__(self, rows, cols, vals, shape):
self.rows = np.asarray(rows, dtype=int)
self.cols = np.asarray(cols, dtype=int)
self.vals = np.asarray(vals, dtype=float)
self.shape = shape
def to_dense(self):
A = np.zeros(self.shape)
for r, c, v in zip(self.rows, self.cols, self.vals):
A[r, c] += v
return A
def to_csr(self):
"""Convert COO to CSR format."""
m = self.shape[0]
# Sort by row
order = np.argsort(self.rows)
rows_s = self.rows[order]
cols_s = self.cols[order]
vals_s = self.vals[order]
# Build indptr: counts per row
indptr = np.zeros(m + 1, dtype=int)
for r in rows_s:
indptr[r + 1] += 1
np.cumsum(indptr, out=indptr)
return SparseMatrixCSR(indptr, cols_s, vals_s, self.shape)
@property
def nnz(self):
return len(self.vals)
class SparseMatrixCSR:
"""
CSR (Compressed Sparse Row) sparse matrix.
Efficient row access and matrix-vector product.
"""
def __init__(self, indptr, col_idx, data, shape):
self.indptr = np.asarray(indptr, dtype=int)
self.col_idx = np.asarray(col_idx, dtype=int)
self.data = np.asarray(data, dtype=float)
self.shape = shape
def matvec(self, x):
"""
Sparse matrix-vector product: y = A @ x
Time: O(nnz) — one pass through non-zeros.
"""
m = self.shape[0]
y = np.zeros(m)
for i in range(m):
start, end = self.indptr[i], self.indptr[i+1]
y[i] = np.dot(self.data[start:end], x[self.col_idx[start:end]])
return y
def to_dense(self):
A = np.zeros(self.shape)
m = self.shape[0]
for i in range(m):
for k in range(self.indptr[i], self.indptr[i+1]):
A[i, self.col_idx[k]] += self.data[k]
return A
@property
def nnz(self):
return len(self.data)
# --- Test ---
rows_t = [0, 0, 1, 2, 2]
cols_t = [0, 2, 1, 0, 2]
vals_t = [1, 2, 3, 4, 5]
shape_t = (3, 3)
coo = SparseMatrixCOO(rows_t, cols_t, vals_t, shape_t)
csr = coo.to_csr()
x_test = np.array([1.0, 2.0, 3.0])
y_sparse = csr.matvec(x_test)
y_dense = coo.to_dense() @ x_test
print('Dense matrix:')
print(coo.to_dense())
print(f'Sparse matvec: {y_sparse}')
print(f'Dense matvec: {y_dense}')
print(f'Match: {np.allclose(y_sparse, y_dense)}')Dense matrix:
[[1. 0. 2.]
[0. 3. 0.]
[4. 0. 5.]]
Sparse matvec: [ 7. 6. 19.]
Dense matvec: [ 7. 6. 19.]
Match: True
6. Experiments¶
# --- Experiment: Sparse vs Dense matvec timing ---
# Hypothesis: sparse matvec is faster when nnz << n^2.
# Try changing: N, NNZ_PER_ROW
import time
N = 1000 # <-- try 500, 2000
NNZ_PER_ROW = 5 # <-- try 3, 10, 50
# Build sparse matrix
rows_e, cols_e, vals_e = [], [], []
for i in range(N):
js = rng.choice(N, NNZ_PER_ROW, replace=False)
for j in js:
rows_e.append(i); cols_e.append(j); vals_e.append(rng.normal())
coo_e = SparseMatrixCOO(rows_e, cols_e, vals_e, (N, N))
csr_e = coo_e.to_csr()
A_dense_e = coo_e.to_dense()
x_e = rng.normal(0, 1, N)
# Time sparse matvec
N_REPS = 20
t0 = time.perf_counter()
for _ in range(N_REPS):
y_s = csr_e.matvec(x_e)
t_sparse = (time.perf_counter() - t0) / N_REPS
# Time dense matvec
t0 = time.perf_counter()
for _ in range(N_REPS):
y_d = A_dense_e @ x_e
t_dense = (time.perf_counter() - t0) / N_REPS
sparsity = 1 - csr_e.nnz / (N**2)
print(f'Matrix: {N}x{N}, {NNZ_PER_ROW} nnz/row')
print(f'Total nnz: {csr_e.nnz} ({100*(1-sparsity):.2f}% non-zero)')
print(f'Sparse matvec: {t_sparse*1000:.3f} ms')
print(f'Dense matvec: {t_dense*1000:.3f} ms')
print(f'Speedup: {t_dense/t_sparse:.1f}x')
print(f'Results match: {np.allclose(y_s, y_d)}')Matrix: 1000x1000, 5 nnz/row
Total nnz: 5000 (0.50% non-zero)
Sparse matvec: 3.362 ms
Dense matvec: 0.471 ms
Speedup: 0.1x
Results match: True
7. Exercises¶
Easy 1. For a tridiagonal matrix (3 nnz per row), compute: total elements, nnz, sparsity fraction, memory ratio (sparse COO vs dense).
Easy 2. Implement SparseMatrixCSR.transpose() that returns a new CSR matrix representing without converting to dense.
Medium 1. Implement sparse matrix addition in CSR format: given two CSR matrices and with the same shape, compute in CSR format.
Medium 2. The power method for finding the dominant eigenvector (ch171) only uses matrix-vector products. Implement it using SparseMatrixCSR.matvec and apply it to a large sparse graph Laplacian to find the Fiedler vector (ch191).
Hard. Implement the conjugate gradient method for solving when is symmetric positive definite and sparse. Each iteration uses only one sparse matvec. Compare convergence rate to dense Gaussian elimination on a 1D Laplacian system of size .
8. Mini Project¶
# --- Mini Project: PageRank via Sparse Matrix-Vector Products ---
# Problem: Compute PageRank scores for a web graph using the power method on a sparse matrix.
# PageRank: r = d * A_norm @ r + (1-d)/n * ones
# where A_norm is the column-normalized adjacency matrix.
n_pages = 30
N_LINKS = 80 # total directed edges
DAMPING = 0.85
# Random web graph
src = rng.integers(0, n_pages, N_LINKS)
dst = rng.integers(0, n_pages, N_LINKS)
# Remove self-loops
mask = src != dst
src, dst = src[mask], dst[mask]
# Build column-normalized adjacency (each column sums to 1)
A_pr = np.zeros((n_pages, n_pages))
for s, d in zip(src, dst):
A_pr[d, s] += 1.0 # column s is source, row d is destination
# Normalize columns
col_sums = A_pr.sum(axis=0)
col_sums[col_sums == 0] = 1 # dangling nodes
A_pr /= col_sums[None, :]
# Power iteration for PageRank
r = np.ones(n_pages) / n_pages
for _ in range(100):
r_new = DAMPING * A_pr @ r + (1 - DAMPING) / n_pages * np.ones(n_pages)
if np.linalg.norm(r_new - r) < 1e-10:
break
r = r_new
print('PageRank scores (top 10 pages):')
top_pages = np.argsort(r)[::-1][:10]
for rank_i, page_i in enumerate(top_pages):
print(f' Rank {rank_i+1}: Page {page_i} score={r[page_i]:.4f}')
fig, axes = plt.subplots(1, 2, figsize=(11, 4))
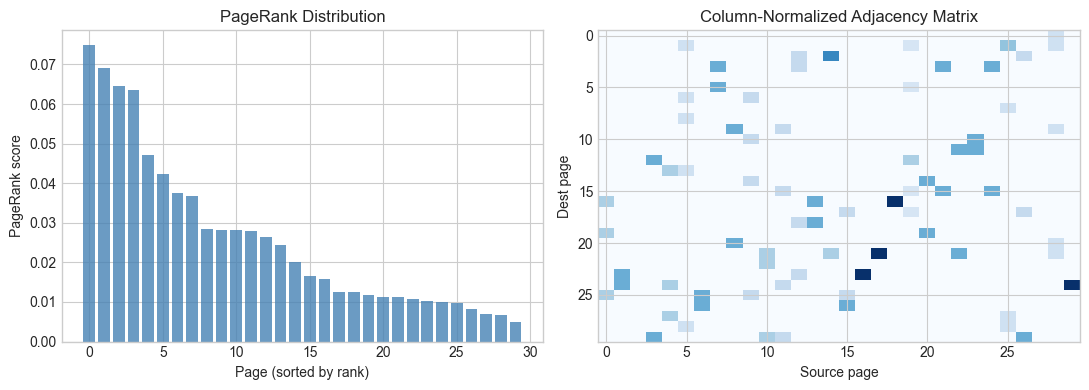
axes[0].bar(range(n_pages), r[np.argsort(r)[::-1]], color='steelblue', alpha=0.8)
axes[0].set_xlabel('Page (sorted by rank)')
axes[0].set_ylabel('PageRank score')
axes[0].set_title('PageRank Distribution')
axes[1].imshow(A_pr, cmap='Blues', aspect='auto')
axes[1].set_title('Column-Normalized Adjacency Matrix')
axes[1].set_xlabel('Source page')
axes[1].set_ylabel('Dest page')
plt.tight_layout()
plt.show()PageRank scores (top 10 pages):
Rank 1: Page 24 score=0.0748
Rank 2: Page 3 score=0.0690
Rank 3: Page 15 score=0.0645
Rank 4: Page 29 score=0.0634
Rank 5: Page 21 score=0.0472
Rank 6: Page 23 score=0.0423
Rank 7: Page 12 score=0.0375
Rank 8: Page 26 score=0.0367
Rank 9: Page 25 score=0.0284
Rank 10: Page 11 score=0.0283
9. Chapter Summary & Connections¶
Sparse matrices store only non-zero entries: COO format stores (row, col, val) triples; CSR stores sorted rows with a pointer array for row access and matrix-vector products.
Sparsity arises structurally in graphs, physics simulations, and text data — where interaction is local or categorical.
The speedup from sparse matvec scales with sparsity: for , computation is orders of magnitude faster than dense.
PageRank, graph spectral methods (ch191), and iterative solvers all use sparse matrix-vector products as their core operation.
Forward: Sparse representations underlie nearly all large-scale linear algebra used in ch191 (Graph Embedding) and ch271–300 (Part IX — Data Science). Gradient computation in deep learning uses sparse Jacobians implicitly. The conjugate gradient method from the Hard exercise is the standard solver for large sparse symmetric positive definite systems encountered in physics and engineering.
Backward: The sparsity pattern of the graph Laplacian (ch191) and the adjacency matrix (ch151) is exactly what makes spectral graph methods computationally feasible at scale.