Prerequisites: ch151 (Matrices), ch154 (Matrix Multiplication), ch172 (Diagonalization), ch173 (SVD) You will learn:
What the Kronecker product computes and how it arises
The vec() operator and the identity vec(AXB) = (B⊤ ⊗ A) vec(X)
Block diagonal and block circulant matrices
Separable filters in signal/image processing
How structure enables orders-of-magnitude speedups Environment: Python 3.x, numpy, matplotlib
1. Concept¶
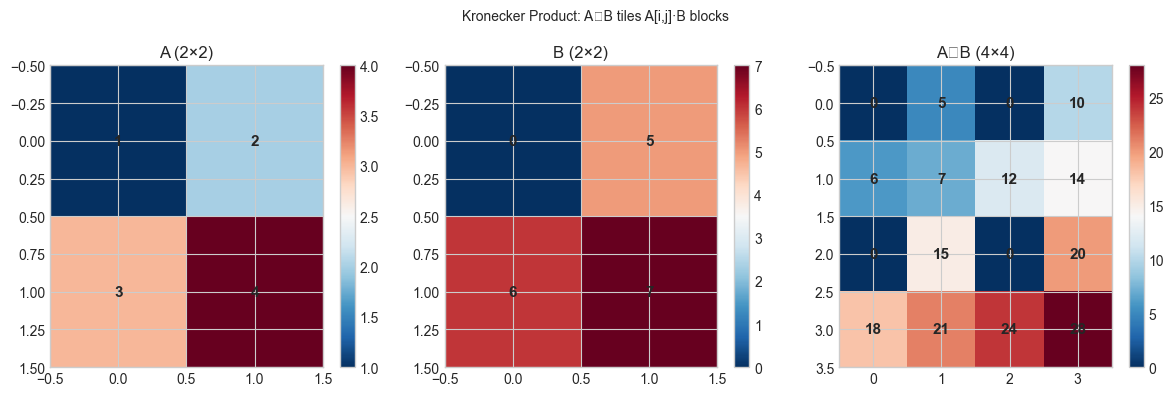
The Kronecker product tiles copies of scaled by entries of to produce a block matrix. If and , then .
Kronecker products arise whenever two independent linear systems act on a combined space — the joint action is the Kronecker product of the individual actions. Examples: 2D image filtering (row filter ⊗ column filter), multi-task learning (task covariance ⊗ feature covariance), and Sylvester matrix equations.
Common misconception: only holds when the shapes are compatible for standard matrix multiplication.
2. Intuition & Mental Models¶
Grid interpretation: if describes row interactions and describes column interactions, describes the combined row-column interaction on a 2D grid — exactly what a separable 2D filter does.
The vec() operator stacks matrix columns into a vector. Key identity:
This converts matrix equations to vector equations — turns (Sylvester equation) into a standard linear system.
Block diagonal matrices appear when independent subsystems share a common interface. Each block can be solved independently: total cost vs for the dense case.
3. Visualization¶
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-v0_8-whitegrid')
rng = np.random.default_rng(42)
def kron(A, B):
"""Kronecker product A ⊗ B."""
m, n = A.shape; p, q = B.shape
C = np.zeros((m*p, n*q))
for i in range(m):
for j in range(n):
C[i*p:(i+1)*p, j*q:(j+1)*q] = A[i,j] * B
return C
A = np.array([[1.,2.],[3.,4.]])
B = np.array([[0.,5.],[6.,7.]])
K = kron(A, B)
fig, axes = plt.subplots(1, 3, figsize=(12, 4))
for ax, mat, title in [(axes[0],A,'A (2×2)'),(axes[1],B,'B (2×2)'),(axes[2],K,'A⊗B (4×4)')]:
im = ax.imshow(mat, cmap='RdBu_r', aspect='auto')
for i in range(mat.shape[0]):
for j in range(mat.shape[1]):
ax.text(j,i,f'{mat[i,j]:.0f}',ha='center',va='center',fontsize=11,fontweight='bold')
ax.set_title(title)
plt.colorbar(im, ax=ax)
plt.suptitle('Kronecker Product: A⊗B tiles A[i,j]·B blocks', fontsize=10)
plt.tight_layout()
plt.show()
print('Verify with numpy:', np.allclose(K, np.kron(A, B)))C:\Users\user\AppData\Local\Temp\ipykernel_19440\2745857543.py:28: UserWarning: Glyph 8855 (\N{CIRCLED TIMES}) missing from font(s) Arial.
plt.tight_layout()
c:\Users\user\OneDrive\Documents\book\.venv\Lib\site-packages\IPython\core\pylabtools.py:170: UserWarning: Glyph 8855 (\N{CIRCLED TIMES}) missing from font(s) Arial.
fig.canvas.print_figure(bytes_io, **kw)
Verify with numpy: True
4. Mathematical Formulation¶
Definition:
Key algebraic properties:
Mixed product:
Transpose:
Inverse: (when both exist)
Eigenvalues: if and , then
vec identity (vectorization):
Sylvester equation : using vec identity becomes .
5. Python Implementation¶
def vec(X):
"""Stack columns of X into a vector (column-major order)."""
return X.T.flatten()
def unvec(v, m, n):
"""Inverse vec: reshape to (m, n) matrix."""
return v.reshape(n, m).T
# Verify vec identity: vec(AXB) = (B.T ⊗ A) @ vec(X)
A_v = rng.normal(0,1,(3,3))
B_v = rng.normal(0,1,(3,3))
X_v = rng.normal(0,1,(3,3))
lhs = vec(A_v @ X_v @ B_v)
rhs = np.kron(B_v.T, A_v) @ vec(X_v)
print(f'vec identity error: {np.linalg.norm(lhs - rhs):.2e} (should be ~0)')
# Solve Sylvester equation AX + XB = C via Kronecker product
def solve_sylvester(A, B, C):
"""
Solve AX + XB = C.
Converts to (I⊗A + B.T⊗I) vec(X) = vec(C).
"""
n = A.shape[0]
I = np.eye(n)
M = np.kron(I, A) + np.kron(B.T, I)
vec_X = np.linalg.solve(M, vec(C))
return unvec(vec_X, n, n)
C_v = rng.normal(0,1,(3,3))
X_sol = solve_sylvester(A_v, B_v, C_v)
resid = np.linalg.norm(A_v @ X_sol + X_sol @ B_v - C_v)
print(f'Sylvester residual: {resid:.2e}')vec identity error: 5.56e-16 (should be ~0)
Sylvester residual: 1.87e-15
6. Experiments¶
# --- Experiment: 2D separable convolution via Kronecker product ---
# Hypothesis: (H_row ⊗ H_col) vec(image) = vec(filtered image)
# Try changing: FILTER_TYPE
FILTER_TYPE = 'smooth' # <-- try 'edge', 'sharpen'
def conv_matrix_1d(h, n):
C = np.zeros((n, n))
pad = len(h) // 2
for i in range(n):
for ki, hv in enumerate(h):
j = i + ki - pad
if 0 <= j < n:
C[i, j] = hv
return C
n_img = 25
image = rng.normal(0,1,(n_img,n_img))
if FILTER_TYPE == 'smooth':
h = np.array([0.25, 0.5, 0.25])
elif FILTER_TYPE == 'edge':
h = np.array([-1., 0., 1.])
else:
h = np.array([-1., 5., -1.])
H = conv_matrix_1d(h, n_img)
H_2d = np.kron(H, H) # separable 2D filter
img_filtered = (H_2d @ image.flatten()).reshape(n_img, n_img)
fig, axes = plt.subplots(1, 2, figsize=(10, 4))
axes[0].imshow(image, cmap='gray')
axes[0].set_title('Original')
axes[1].imshow(img_filtered, cmap='gray')
axes[1].set_title(f'{FILTER_TYPE} filter via H_row ⊗ H_col')
plt.tight_layout()
plt.show()
print(f'2D filter applied via single Kronecker product: {H_2d.shape} matrix')C:\Users\user\AppData\Local\Temp\ipykernel_19440\3127294582.py:36: UserWarning: Glyph 8855 (\N{CIRCLED TIMES}) missing from font(s) Arial.
plt.tight_layout()
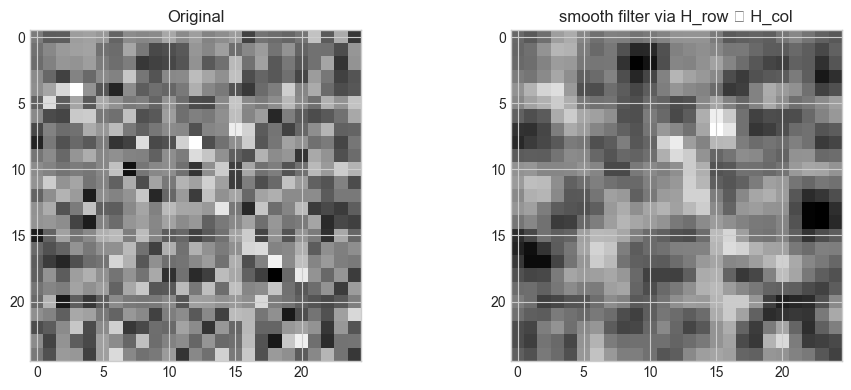
2D filter applied via single Kronecker product: (625, 625) matrix
7. Exercises¶
Easy 1. Compute for a identity and arbitrary . What is the pattern? What is ?
Easy 2. Verify the eigenvalue property: has eigenvalues where are eigenvalues of and of . Test with two symmetric matrices.
Medium 1. Implement block-diagonal matrix-vector multiply: given matrices and a vector partitioned accordingly, compute without forming the full block-diagonal matrix.
Medium 2. The Kronecker-factored least squares problem: minimize over . Use the vec identity to reduce it to a standard least-squares problem and solve.
Hard. Build a 3D Laplacian via Kronecker products: where is the 1D Laplacian. Solve on a small 3D grid using the CG method from ch196.
8. Mini Project¶
# --- Mini Project: Multi-Dimensional Separable Regression ---
# Fit f(x,y) on a 2D grid using Kronecker-structured design matrix.
NX, NY, N_OBS = 12, 10, 150
a_true = np.sin(np.linspace(0, np.pi, NX))
b_true = np.cos(np.linspace(0, np.pi, NY))
F_true = np.outer(a_true, b_true)
obs_x = rng.integers(0, NX, N_OBS)
obs_y = rng.integers(0, NY, N_OBS)
y_obs = F_true[obs_x, obs_y] + 0.05 * rng.normal(0,1,N_OBS)
# Design matrix: each row is e_x ⊗ e_y (one-hot for grid point)
A_design = np.zeros((N_OBS, NX * NY))
for k, (xi, yi) in enumerate(zip(obs_x, obs_y)):
A_design[k, xi * NY + yi] = 1.0
F_hat = np.linalg.lstsq(A_design, y_obs, rcond=None)[0].reshape(NX, NY)
err = np.linalg.norm(F_hat - F_true, 'fro')
fig, axes = plt.subplots(1,2,figsize=(10,4))
axes[0].imshow(F_true, cmap='RdBu_r')
axes[0].set_title('True f(x,y) = sin(x)·cos(y)')
axes[1].imshow(F_hat, cmap='RdBu_r')
axes[1].set_title(f'Estimated (err={err:.4f})')
plt.tight_layout()
plt.show()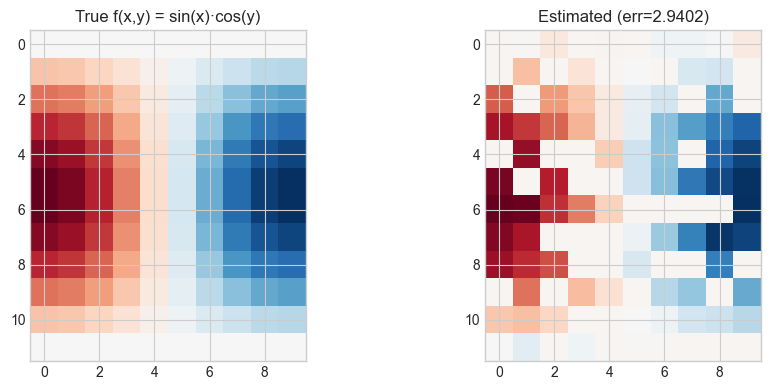
9. Chapter Summary & Connections¶
The Kronecker product encodes the joint action of two independent linear maps; it arises in 2D filtering, multi-task learning, and matrix equations (ch154).
The vec identity converts matrix equations to vector equations, enabling Kronecker-based solvers.
Separable structure reduces 2D filter application from to ; 3D reductions are even more dramatic.
Block diagonal matrices arise in independent subsystems and can be solved block-by-block.
Forward: Kronecker structure appears in multi-task Gaussian process regression, tensor decompositions (used in recommender systems at scale), and K-FAC (Kronecker-Factored Approximate Curvature) for efficient neural network optimization. The 3D Laplacian via Kronecker products connects to physics simulation.
Backward: This generalizes ch154 (Matrix Multiplication) to block-structured settings and applies ch172 (Diagonalization) — the eigenvalue product property follows directly from diagonalizing both and simultaneously.