Prerequisites: ch173 (SVD), ch174 (PCA), ch192 (Rank), ch194 (Condition Numbers) You will learn:
The Johnson-Lindenstrauss lemma and random projections
Randomized SVD: fast low-rank approximation with provable error bounds
Sketch-and-solve for large least-squares problems
When randomized algorithms are preferable to deterministic ones
Error bounds and the role of oversampling Environment: Python 3.x, numpy, matplotlib
1. Concept¶
Full SVD of an matrix costs — infeasible for . Randomized algorithms exploit approximate low-rank structure: a random projection onto a small subspace captures most of the information, then we solve exactly in that reduced space.
These are not heuristics — they have provable error bounds. The Johnson-Lindenstrauss (JL) lemma guarantees that a random projection to dimensions approximately preserves all pairwise distances simultaneously.
Common misconception: randomized methods are Monte Carlo algorithms that give different answers each run. In practice, a single random draw is taken; subsequent computation is exact. The randomness is in the construction of the projection, not the solution.
2. Intuition & Mental Models¶
Random projection: imagine points in . Project onto a random -dimensional subspace. Since the subspace is random, it is unlikely to be aligned with any particular structured direction — so distances are approximately preserved. The JL lemma makes this precise.
Randomized SVD: (1) multiply by a random matrix to sketch the column space, (2) find an orthonormal basis for the sketch, (3) project onto and compute the SVD of the small . This replaces with where .
Think of it as: casting a random fishing net in the high-dimensional space and catching the most important directions — then rigorously computing in the smaller space spanned by the catch.
3. Visualization¶
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-v0_8-whitegrid')
rng = np.random.default_rng(42)
# JL Lemma demo: random projection preserves pairwise distances
N_PTS, D_ORIG, K_PROJ = 300, 500, 40
X = rng.normal(0, 1, (N_PTS, D_ORIG))
Omega = rng.normal(0, 1/np.sqrt(K_PROJ), (D_ORIG, K_PROJ))
X_proj = X @ Omega
# Sample pairwise distances
N_PAIRS = 1000
i_idx = rng.integers(0, N_PTS, N_PAIRS)
j_idx = rng.integers(0, N_PTS, N_PAIRS)
mask = i_idx != j_idx
i_idx, j_idx = i_idx[mask], j_idx[mask]
dists_orig = np.linalg.norm(X[i_idx] - X[j_idx], axis=1)
dists_proj = np.linalg.norm(X_proj[i_idx] - X_proj[j_idx], axis=1)
ratios = dists_proj / (dists_orig + 1e-10)
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
axes[0].scatter(dists_orig, dists_proj, alpha=0.2, s=8, color='steelblue')
lim = max(dists_orig.max(), dists_proj.max())
axes[0].plot([0, lim], [0, lim], 'r--', lw=2, label='Perfect preservation')
axes[0].set_xlabel('Original distance')
axes[0].set_ylabel('Projected distance')
axes[0].set_title(f'JL Lemma: {D_ORIG}D → {K_PROJ}D')
axes[0].legend(fontsize=8)
axes[1].hist(ratios, bins=40, color='steelblue', alpha=0.8)
axes[1].axvline(1.0, color='crimson', lw=2)
axes[1].set_xlabel('dist_proj / dist_orig')
axes[1].set_ylabel('Count')
axes[1].set_title(f'Distance ratio (mean={ratios.mean():.3f}, std={ratios.std():.3f})')
plt.tight_layout()
plt.show()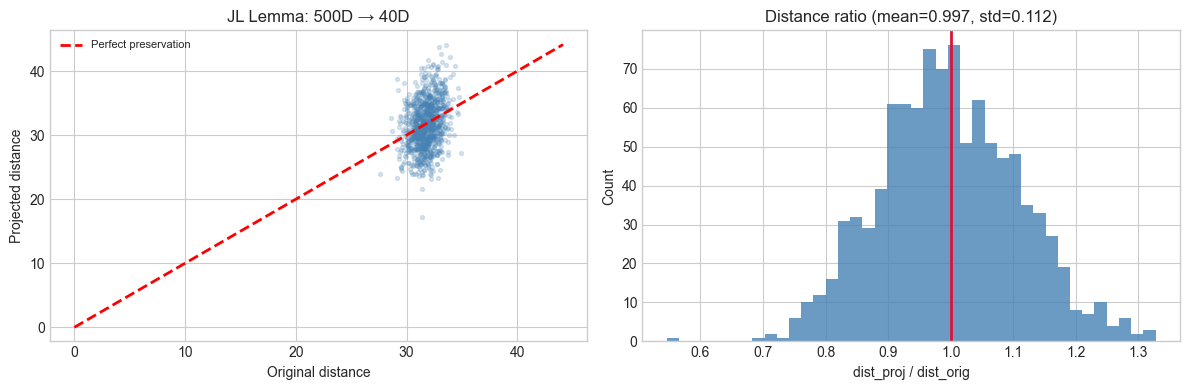
4. Mathematical Formulation¶
JL Lemma: for any and points in , a random linear map with satisfies with high probability:
Randomized SVD (Halko, Martinsson, Tropp 2011):
random Gaussian, = oversampling (typically 5–10)
, then (QR decomposition)
, then (exact SVD of small matrix)
: final answer
Error bound:
5. Python Implementation¶
def randomized_svd(A, k, oversampling=10, power_iter=2, rng=rng):
"""
Randomized SVD: approximate top-k singular values/vectors.
Args:
A: (m, n) matrix
k: target rank
oversampling: extra columns for stability (p in theory)
power_iter: subspace iteration steps (improves accuracy)
Returns:
U: (m, k), s: (k,), Vt: (k, n)
"""
m, n = A.shape
rank = k + oversampling
# Random sketch
Omega = rng.standard_normal((n, rank))
Y = A @ Omega
# Power iteration: improves accuracy for slowly-decaying spectra
for _ in range(power_iter):
Y = A @ (A.T @ Y)
# Orthonormal basis for range of Y
Q, _ = np.linalg.qr(Y)
Q = Q[:, :rank]
# Project and compute small SVD
B = Q.T @ A
U_hat, s, Vt = np.linalg.svd(B, full_matrices=False)
U = Q @ U_hat
return U[:, :k], s[:k], Vt[:k, :]
# Validate on a known low-rank matrix
M, N, K_true = 400, 300, 15
U_t = rng.normal(0,1,(M,K_true)); V_t = rng.normal(0,1,(N,K_true))
A_lr = U_t @ V_t.T + 0.01 * rng.normal(0,1,(M,N))
U_ex, s_ex, Vt_ex = np.linalg.svd(A_lr, full_matrices=False)
for k_test in [5, 10, 15, 20]:
U_r, s_r, Vt_r = randomized_svd(A_lr, k_test)
A_k_ex = U_ex[:,:k_test] @ np.diag(s_ex[:k_test]) @ Vt_ex[:k_test]
A_k_r = U_r @ np.diag(s_r) @ Vt_r
err_ex = np.linalg.norm(A_lr - A_k_ex, 'fro')
err_r = np.linalg.norm(A_lr - A_k_r, 'fro')
print(f'k={k_test}: exact={err_ex:.4f}, rand={err_r:.4f}, ratio={err_r/err_ex:.3f}')k=5: exact=994.8164, rand=994.8164, ratio=1.000
k=10: exact=642.5870, rand=642.5870, ratio=1.000
k=15: exact=3.3028, rand=3.3028, ratio=1.000
k=20: exact=3.2051, rand=3.2181, ratio=1.004
6. Experiments¶
# --- Experiment: Randomized SVD speed vs accuracy ---
# Hypothesis: randomized SVD is faster with small accuracy loss for low-rank matrices.
# Try changing: M_exp, N_exp, K_exp, oversampling
import time
M_exp, N_exp, K_exp = 800, 600, 12
U_e = rng.normal(0,1,(M_exp,K_exp)); V_e = rng.normal(0,1,(N_exp,K_exp))
A_exp = U_e @ V_e.T + 0.01 * rng.normal(0,1,(M_exp,N_exp))
t0 = time.perf_counter()
U_full, s_full, Vt_full = np.linalg.svd(A_exp, full_matrices=False)
t_full = time.perf_counter() - t0
print(f'Full SVD: {t_full*1000:.1f}ms')
for k_r in [5, 10, 15, 20]:
t0 = time.perf_counter()
_, s_r, _ = randomized_svd(A_exp, k_r)
t_r = time.perf_counter() - t0
sv_err = np.linalg.norm(s_r - s_full[:k_r]) / np.linalg.norm(s_full[:k_r])
print(f' Rand k={k_r}: {t_r*1000:.1f}ms, SV err={sv_err:.4f}')Full SVD: 241.9ms
Rand k=5: 5.7ms, SV err=0.0000
Rand k=10: 7.1ms, SV err=0.0000
Rand k=15: 11.0ms, SV err=0.0001
Rand k=20: 12.9ms, SV err=0.0001
7. Exercises¶
Easy 1. Implement the JL projection and measure mean distance ratio for points in dimensions projected to . How does the ratio’s standard deviation change with ?
Easy 2. How does the oversampling parameter affect accuracy in randomized SVD? Fix , vary , plot Frobenius error vs .
Medium 1. Implement sketch-and-solve for overdetermined least squares: given with , form random Gaussian (), solve the sketched system. Compare accuracy and timing to the exact pseudoinverse solution.
Medium 2. The power iteration step improves accuracy for matrices with slowly-decaying singular values. Demonstrate this: construct a matrix with (slow decay), run randomized SVD with power_iter=0,1,2,3, plot Frobenius error vs number of power iteration steps.
Hard. Implement streaming randomized SVD: process in row-blocks of size , accumulating the sketch without storing the full matrix. Verify the result matches batch randomized SVD.
8. Mini Project¶
# --- Mini Project: Fast Image Compression via Randomized SVD ---
import time
H, W, K_img = 150, 200, 10
U_img = rng.normal(0,1,(H,K_img)); V_img = rng.normal(0,1,(W,K_img))
img = rng.normal(0,1,(H,W)) * 0.1 + (U_img @ V_img.T)
img = (img - img.min()) / (img.max() - img.min())
K_VALS = [5, 10, 15, 25]
fig, axes = plt.subplots(2, len(K_VALS)+1, figsize=(14, 5))
axes[0,0].imshow(img, cmap='gray'); axes[0,0].set_title('Original'); axes[0,0].axis('off')
axes[1,0].axis('off')
for col, k_c in enumerate(K_VALS):
# Randomized
t0 = time.perf_counter()
Ur, sr, Vtr = randomized_svd(img, k_c)
t_r = time.perf_counter() - t0
img_r = np.clip(Ur @ np.diag(sr) @ Vtr, 0, 1)
err_r = np.linalg.norm(img - img_r, 'fro')
# Exact
t0 = time.perf_counter()
Ue, se, Vte = np.linalg.svd(img, full_matrices=False)
t_e = time.perf_counter() - t0
img_e = np.clip(Ue[:,:k_c] @ np.diag(se[:k_c]) @ Vte[:k_c], 0, 1)
err_e = np.linalg.norm(img - img_e, 'fro')
axes[0,col+1].imshow(img_r, cmap='gray')
axes[0,col+1].set_title(f'Rand k={k_c}\n{t_r*1000:.0f}ms err={err_r:.3f}', fontsize=7)
axes[0,col+1].axis('off')
axes[1,col+1].imshow(img_e, cmap='gray')
axes[1,col+1].set_title(f'Exact k={k_c}\n{t_e*1000:.0f}ms err={err_e:.3f}', fontsize=7)
axes[1,col+1].axis('off')
fig.suptitle('Image Compression: Randomized (top) vs Exact SVD (bottom)', fontsize=10)
plt.tight_layout()
plt.show()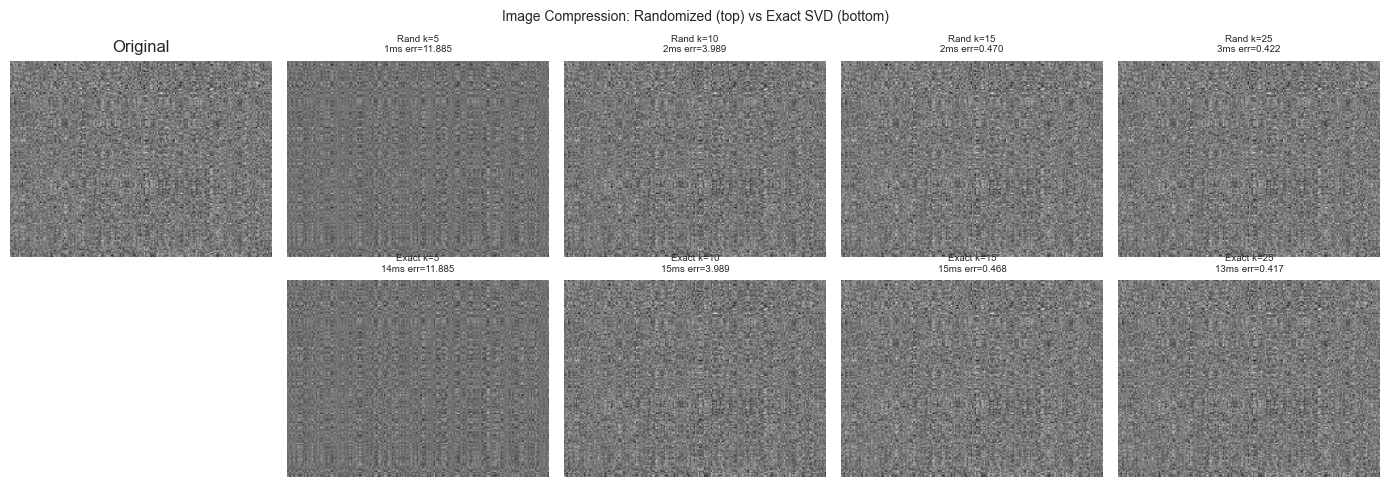
9. Chapter Summary & Connections¶
Randomized algorithms project onto random low-dimensional subspaces, then solve exactly — achieving near-optimal accuracy with provable guarantees and dramatic speedups (ch173).
JL lemma: dimensions suffice to approximately preserve pairwise distances. Foundational to approximate nearest-neighbor search.
Randomized SVD replaces with ; power iteration improves accuracy for slowly-decaying spectra.
Sketch-and-solve extends these ideas to least-squares problems.
Forward: Randomized methods enable the SVD-based algorithms of ch180–191 at dataset scales encountered in Part IX (millions of samples, high-dimensional features). The JL lemma underpins locality-sensitive hashing used in ch289 (Collaborative Filtering at Scale). Random projections also appear in compressed sensing and sparse recovery.
Backward: This chapter extends ch173 (SVD), ch174 (PCA), and ch180 (Image Compression) to the large-scale regime. The error bounds rely on probability theory that will be formalized in Part VIII (ch241–270).