(Uses logarithms from ch043; probability distributions from ch248; connects directly to ML loss functions)
1. The Central Question¶
How do you measure the amount of information in a message?
Claude Shannon (1948) answered this by connecting information to surprise. A rare event carries more information than a common one. If a fair coin lands heads, you learn 1 bit. If a 1-in-a-million event occurs, you learn ~20 bits.
The self-information (surprisal) of event with probability :
Or in nats (natural units, used in ML):
2. Why This Definition?¶
Three axioms uniquely determine this form:
is a continuous, decreasing function of
— certain events carry no information
— independent events add information
The only function satisfying all three: .
import numpy as np
import matplotlib.pyplot as plt
def self_information(p: float | np.ndarray, base: str = 'nats') -> float | np.ndarray:
"""Surprisal: -log(p). base='bits' uses log2, 'nats' uses ln."""
if base == 'bits':
return -np.log2(np.clip(p, 1e-15, 1.0))
return -np.log(np.clip(p, 1e-15, 1.0))
p_vals = np.linspace(0.001, 1.0, 500)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
ax1.plot(p_vals, self_information(p_vals, 'bits'), 'b-', lw=2)
ax1.scatter([0.5, 0.1, 0.01, 0.001],
self_information(np.array([0.5, 0.1, 0.01, 0.001]), 'bits'),
s=80, color='red', zorder=5)
for p, label in [(0.5, 'fair coin\n(1 bit)'),
(0.1, '10% event\n(3.3 bits)')]:
ax1.annotate(label, xy=(p, self_information(p, 'bits')),
xytext=(p+0.1, self_information(p, 'bits')+1), fontsize=8,
arrowprops=dict(arrowstyle='->', lw=1))
ax1.set_xlabel('Probability p'); ax1.set_ylabel('Self-information (bits)')
ax1.set_title('Surprisal = -log₂(p)')
# Verify additivity for independent events
p1, p2 = 0.3, 0.4
ax2.bar(['I(p1)', 'I(p2)', 'I(p1)+I(p2)', 'I(p1·p2)'],
[self_information(p1, 'bits'),
self_information(p2, 'bits'),
self_information(p1, 'bits') + self_information(p2, 'bits'),
self_information(p1*p2, 'bits')],
color=['steelblue','coral','mediumpurple','mediumpurple'],
edgecolor='white')
ax2.set_title('Additivity: I(p1·p2) = I(p1) + I(p2)')
ax2.set_ylabel('bits')
plt.tight_layout()
plt.show()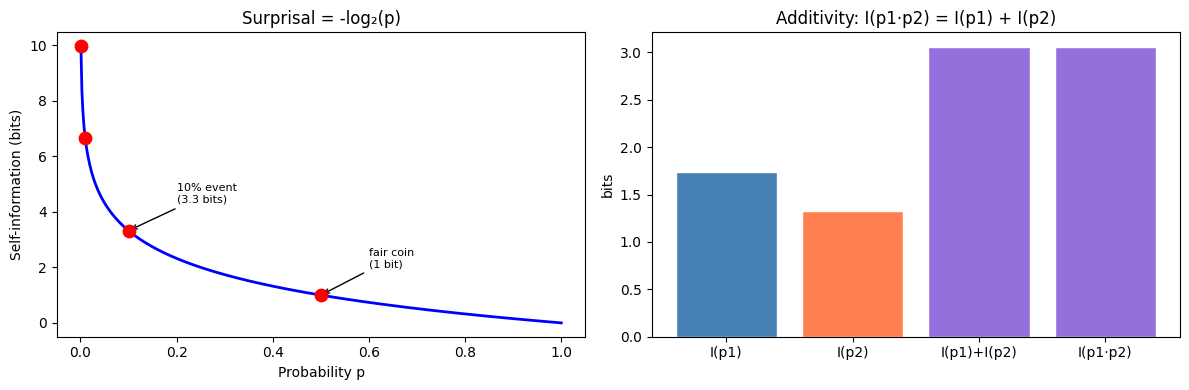
3. Cross-Entropy — The ML Loss Function¶
Cross-entropy measures the average number of bits needed to encode events from distribution using a code designed for distribution :
In classification, = true labels (one-hot), = model predictions. Minimizing cross-entropy is identical to maximum likelihood estimation.
def cross_entropy(
p_true: np.ndarray,
q_pred: np.ndarray,
eps: float = 1e-15,
) -> float:
"""Cross-entropy H(P, Q) in nats. p_true and q_pred are 1D probability vectors."""
return -np.sum(p_true * np.log(np.clip(q_pred, eps, 1.0)))
def binary_cross_entropy(y_true: np.ndarray, y_pred: np.ndarray) -> float:
"""Binary cross-entropy per sample, averaged."""
eps = 1e-15
y_pred = np.clip(y_pred, eps, 1 - eps)
return -np.mean(y_true * np.log(y_pred) + (1 - y_true) * np.log(1 - y_pred))
# Effect of prediction confidence on CE loss
y_true_probs = np.array([1.0, 0.0, 0.0]) # true class = 0
predictions = [
(np.array([0.9, 0.05, 0.05]), 'Confident correct'),
(np.array([0.5, 0.3, 0.2]), 'Uncertain correct'),
(np.array([0.33, 0.34, 0.33]), 'Uniform (max entropy)'),
(np.array([0.1, 0.5, 0.4]), 'Uncertain wrong'),
(np.array([0.01, 0.9, 0.09]), 'Confident wrong'),
]
print(f"True class: [1, 0, 0]")
print(f"{'Prediction':<30} {'CE loss':>10}")
print("-" * 44)
for q, label in predictions:
ce = cross_entropy(y_true_probs, q)
print(f"{label:<30} {ce:>10.4f}")
print()
# Binary CE curve
y_pred_range = np.linspace(0.001, 0.999, 300)
ce_true_1 = -np.log(y_pred_range) # y=1: loss = -log(p)
ce_true_0 = -np.log(1 - y_pred_range) # y=0: loss = -log(1-p)
fig, ax = plt.subplots(figsize=(9, 4))
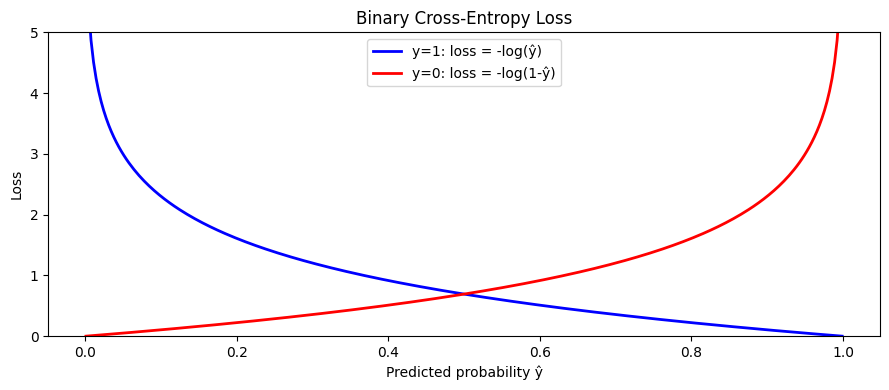
ax.plot(y_pred_range, ce_true_1, 'b-', lw=2, label='y=1: loss = -log(ŷ)')
ax.plot(y_pred_range, ce_true_0, 'r-', lw=2, label='y=0: loss = -log(1-ŷ)')
ax.set_xlabel('Predicted probability ŷ')
ax.set_ylabel('Loss')
ax.set_ylim(0, 5)
ax.set_title('Binary Cross-Entropy Loss')
ax.legend()
plt.tight_layout()
plt.show()True class: [1, 0, 0]
Prediction CE loss
--------------------------------------------
Confident correct 0.1054
Uncertain correct 0.6931
Uniform (max entropy) 1.1087
Uncertain wrong 2.3026
Confident wrong 4.6052
4. What Comes Next¶
Cross-entropy decomposes into entropy plus KL divergence: . ch288 — Entropy develops the entropy term; ch289 — KL Divergence develops the divergence term. Understanding both is necessary to interpret what a neural network’s loss function is actually minimizing.