1. The overfitting problem in deep networks¶
Deep networks have enough capacity to memorise training data entirely — achieving zero training loss while failing on new data (bias–variance tradeoff: ch276). Regularisation techniques constrain the model to generalise.
2. L2 and L1 regularisation¶
Add a penalty to the loss:
L2 (weight decay) shrinks all weights uniformly. L1 pushes small weights to exactly zero, producing sparse solutions. For neural networks, L2 is standard — it corresponds to a Gaussian prior on weights (Bayesian view: ch292).
The L2 gradient update becomes:
import numpy as np
import matplotlib.pyplot as plt
class Dropout:
"""Inverted dropout: scales at training time so inference requires no change."""
def __init__(self, p: float):
"""p = probability of KEEPING a unit (keep_prob). Drop prob = 1-p."""
assert 0 < p <= 1, "p must be in (0, 1]"
self.p = p
self.mask = None
def forward(self, A: np.ndarray, training: bool = True) -> np.ndarray:
if not training or self.p == 1.0:
return A
# Inverted dropout: scale by 1/p so expected value is preserved
self.mask = (np.random.rand(*A.shape) < self.p) / self.p
return A * self.mask
def backward(self, dA: np.ndarray) -> np.ndarray:
return dA * self.mask
# Demonstrate: effect of dropout on activation distribution and co-adaptation
rng = np.random.default_rng(42)
n_neurons, batch = 200, 500
A = rng.normal(0, 1, (n_neurons, batch))
fig, axes = plt.subplots(1, 3, figsize=(13, 4))
p_values = [1.0, 0.8, 0.5]
titles = ['No dropout', 'Dropout p=0.8 (20% dropped)', 'Dropout p=0.5 (50% dropped)']
colors = ['#2ecc71', '#3498db', '#e74c3c']
for ax, p, title, color in zip(axes, p_values, titles, colors):
dp = Dropout(p)
A_out = dp.forward(A, training=True)
# Show neuron activation rates
active_fraction = (A_out != 0).mean(axis=1)
ax.hist(A_out.ravel(), bins=60, color=color, alpha=0.7, density=True, label='Activations')
ax.set_title(f'{title}\nmean={A_out.mean():.3f}, std={A_out.std():.3f}')
ax.set_xlabel('Activation value')
if p < 1.0:
frac = (A_out.ravel() == 0).mean()
ax.text(0.05, 0.9, f'{frac:.0%} zeros', transform=ax.transAxes, fontsize=10,
color='black', bbox=dict(boxstyle='round', facecolor='white', alpha=0.7))
axes[0].set_ylabel('Density')
plt.suptitle('Inverted dropout: mean preserved despite random zeroing', fontsize=11)
plt.tight_layout()
plt.savefig('ch311_dropout.png', dpi=120)
plt.show()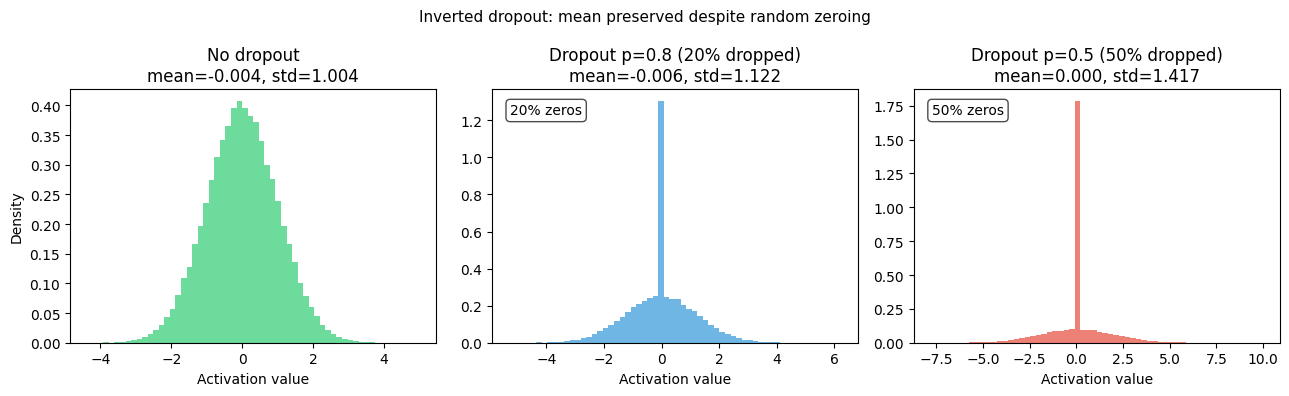
# Train a small network with/without dropout on noisy data
# and compare generalisation
import numpy as np
rng = np.random.default_rng(1)
n_train, n_test = 100, 500
# Noisy 1D regression: y = sin(x) + noise
def make_data(n, noise=0.5):
x = rng.uniform(-np.pi, np.pi, n)
y = np.sin(x) + rng.normal(0, noise, n)
return x[:, None], y[:, None]
X_tr, y_tr = make_data(n_train)
X_te, y_te = make_data(n_test, noise=0.0)
def sigmoid(z): return 1/(1+np.exp(-z))
def sigmoid_grad(z): s=sigmoid(z); return s*(1-s)
def relu(z): return np.maximum(0,z)
def relu_grad(z): return (z>0).astype(float)
def train_net(use_dropout=False, p_keep=0.7, lr=0.01, n_epochs=500, l2=0.001):
rng2 = np.random.default_rng(42)
W1 = rng2.normal(0, np.sqrt(2/1), (64, 1))
b1 = np.zeros((64,1))
W2 = rng2.normal(0, np.sqrt(2/64), (64, 64))
b2 = np.zeros((64,1))
W3 = rng2.normal(0, np.sqrt(2/64), (1, 64))
b3 = np.zeros((1,1))
train_losses, test_losses = [], []
dp1 = Dropout(p_keep)
dp2 = Dropout(p_keep)
for epoch in range(n_epochs):
X = X_tr.T; y = y_tr.T
Z1 = W1@X + b1; A1 = relu(Z1)
if use_dropout: A1 = dp1.forward(A1, training=True)
Z2 = W2@A1 + b2; A2 = relu(Z2)
if use_dropout: A2 = dp2.forward(A2, training=True)
Z3 = W3@A2 + b3; yhat = Z3
loss_train = float(np.mean((yhat - y)**2))
train_losses.append(loss_train)
# Test (no dropout)
A1t = relu(W1@X_te.T + b1)
A2t = relu(W2@A1t + b2)
yhat_t = W3@A2t + b3
test_losses.append(float(np.mean((yhat_t - y_te.T)**2)))
# Backward
B = X.shape[1]
dZ3 = (yhat - y) * 2 / B
dW3 = dZ3 @ A2.T + l2*W3
db3 = dZ3.sum(axis=1, keepdims=True)
dA2 = W3.T @ dZ3
if use_dropout: dA2 = dp2.backward(dA2)
dZ2 = dA2 * relu_grad(Z2)
dW2 = dZ2 @ A1.T + l2*W2
db2 = dZ2.sum(axis=1, keepdims=True)
dA1 = W2.T @ dZ2
if use_dropout: dA1 = dp1.backward(dA1)
dZ1 = dA1 * relu_grad(Z1)
dW1 = dZ1 @ X.T + l2*W1
db1 = dZ1.sum(axis=1, keepdims=True)
for W,dW in [(W1,dW1),(W2,dW2),(W3,dW3)]:
W -= lr*dW
for b,db in [(b1,db1),(b2,db2),(b3,db3)]:
b -= lr*db
return train_losses, test_losses
tr_no, te_no = train_net(use_dropout=False)
tr_do, te_do = train_net(use_dropout=True, p_keep=0.7)
import matplotlib.pyplot as plt
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(te_no, label='Test loss (no dropout)', color='#e74c3c', lw=2)
ax.plot(te_do, label='Test loss (dropout p=0.7)', color='#3498db', lw=2)
ax.plot(tr_no, '--', label='Train loss (no dropout)', color='#e74c3c', alpha=0.5)
ax.plot(tr_do, '--', label='Train loss (dropout p=0.7)', color='#3498db', alpha=0.5)
ax.set_xlabel('Epoch')
ax.set_ylabel('MSE')
ax.set_title('Dropout regularisation: train/test gap with and without')
ax.legend()
ax.set_ylim(0, 1.5)
plt.tight_layout()
plt.savefig('ch311_dropout_effect.png', dpi=120)
plt.show()
print(f"Final test MSE — no dropout: {te_no[-1]:.4f} | dropout: {te_do[-1]:.4f}")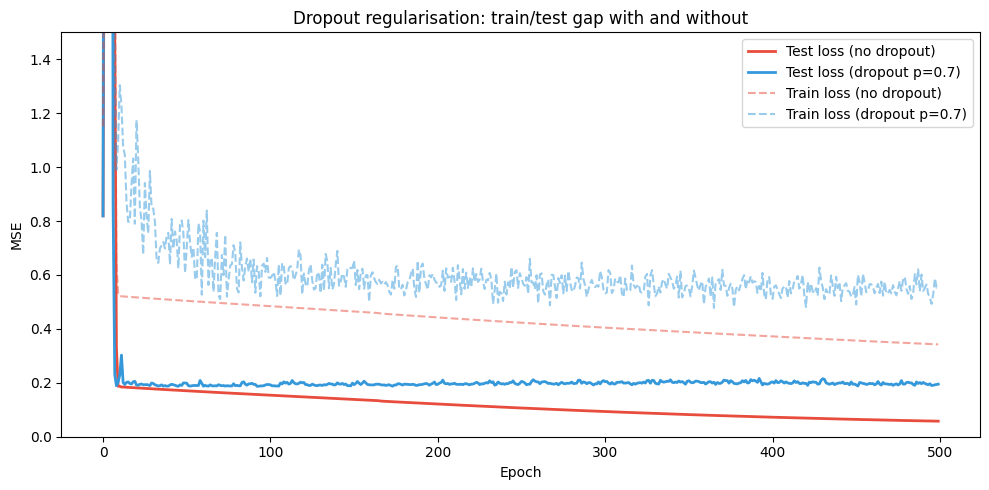
Final test MSE — no dropout: 0.0576 | dropout: 0.1950
3. Other regularisation techniques¶
| Technique | Mechanism | When to use |
|---|---|---|
| L2 (weight decay) | Penalise large weights | Always; cheap to add |
| L1 | Penalise non-zero weights | When sparse features desirable |
| Dropout | Random zeroing during training | Wide MLPs, Transformers |
| DropConnect | Drop weights not activations | Rare; slightly stronger than dropout |
| Data augmentation | Expand training set artificially | Computer vision (ch335) |
| Early stopping | Halt when val loss stops decreasing | Always a good baseline |
| Batch norm | Slight noise from batch statistics | CNNs (not a primary regulariser) |
4. Ensemble interpretation of dropout¶
At test time, dropout approximates an ensemble average over thinned networks (where is the number of dropout units). This is why it reduces variance.
5. Summary¶
L2 regularisation = Gaussian prior on weights; L1 = Laplace prior → sparsity.
Dropout randomly zeroes neurons during training; inverted dropout preserves expected activation.
Regularisation reduces the train-test gap by preventing co-adaptation of neurons.
Use weight decay always; add dropout for wide networks and Transformers.
6. Forward and backward references¶
Used here: overfitting and bias–variance (ch276), gradient update (ch307), Bayesian prior view (ch292).
This will reappear in ch322 — Transformers, where dropout is applied after every attention and feed-forward sublayer, and in ch334 — Project: Neural Net from Scratch.